L-Glutamine (PAMDB000163)
| Record Information | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Version | 1.0 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Update Date | 1/22/2018 12:54:54 PM | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Metabolite ID | PAMDB000163 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Identification | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
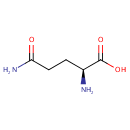
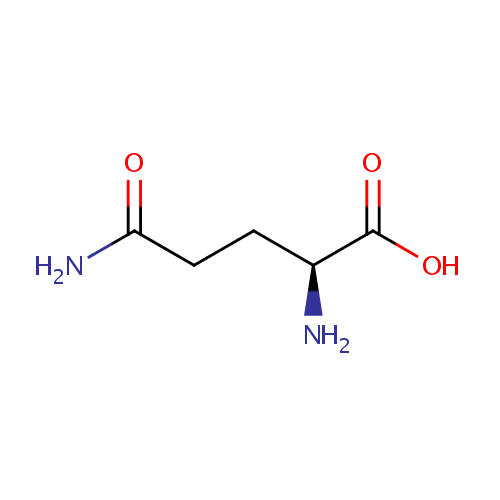
| Name: | L-Glutamine | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Description: | Glutamine (Gln) is one of the 20 amino acids encoded by the standard genetic code. Its side chain is an amide; it is formed by replacing a side-chain hydroxyl of glutamic acid with an amine functional group. Glutamine has a variety of biochemical functions including: 1. As any other amino acid, a major role in protein synthesis. 2. Cellular energy source, next to glucose. 3. Nitrogen donor for many anabolic processes. 4.Carbon source, refilling the Citric acid cycle. (Wikipedia) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Structure | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Synonyms: |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Chemical Formula: | C5H10N2O3 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Average Molecular Weight: | 146.1445 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Monoisotopic Molecular Weight: | 146.069142196 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| InChI Key: | ZDXPYRJPNDTMRX-VKHMYHEASA-N | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| InChI: | InChI=1S/C5H10N2O3/c6-3(5(9)10)1-2-4(7)8/h3H,1-2,6H2,(H2,7,8)(H,9,10)/t3-/m0/s1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| CAS number: | 56-85-9 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| IUPAC Name: | (2S)-2-amino-4-carbamoylbutanoic acid | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Traditional IUPAC Name: | L-glutamine | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| SMILES: | N[C@@H](CCC(N)=O)C(O)=O | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Chemical Taxonomy | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Taxonomy Description | This compound belongs to the class of organic compounds known as l-alpha-amino acids. These are alpha amino acids which have the L-configuration of the alpha-carbon atom. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Kingdom | Organic compounds | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Super Class | Organic acids and derivatives | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Class | Carboxylic acids and derivatives | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Sub Class | Amino acids, peptides, and analogues | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Direct Parent | L-alpha-amino acids | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Alternative Parents | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Substituents |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Molecular Framework | Aliphatic acyclic compounds | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| External Descriptors |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Physical Properties | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| State: | Solid | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Charge: | 0 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Melting point: | 185 °C | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Experimental Properties: |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Predicted Properties |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Biological Properties | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Cellular Locations: | Cytoplasm | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Reactions: | 2 Adenosine triphosphate + L-Glutamine + Water + Hydrogen carbonate >2 ADP + Carbamoylphosphate + L-Glutamate +2 Hydrogen ion + Phosphate Adenosine triphosphate + Water + L-Glutamine > ADP + L-Glutamine + Hydrogen ion + Phosphate Adenosine triphosphate + Water + L-Glutamine > ADP + L-Glutamine + Hydrogen ion + Phosphate Chorismate + L-Glutamine <> 2-Aminobenzoic acid + L-Glutamate + Hydrogen ion + Pyruvic acid L-Glutamine + Water > L-Glutamate + Ammonium L-Glutamine + Phosphoribulosylformimino-AICAR-P > Phosphoribosyl formamidocarboxamide + D-Erythro-imidazole-glycerol-phosphate + L-Glutamate + Hydrogen ion Chorismate + L-Glutamine <> 4-Amino-4-deoxychorismate + L-Glutamate Adenosine triphosphate + L-Glutamate + Ammonium > ADP + L-Glutamine + Hydrogen ion + Phosphate L-Aspartic acid + Adenosine triphosphate + L-Glutamine + Water > Adenosine monophosphate + L-Asparagine + L-Glutamate + Hydrogen ion + Pyrophosphate Adenosine triphosphate + L-Glutamine + tRNA(Gln) > Adenosine monophosphate + L-Glutaminyl-tRNA(Gln) + Pyrophosphate Adenosine triphosphate + L-Glutamine + Water + Xanthylic acid > Adenosine monophosphate + L-Glutamate + Guanosine monophosphate +2 Hydrogen ion + Pyrophosphate Adenosine triphosphate + 5'-Phosphoribosyl-N-formylglycineamide + L-Glutamine + Water <> ADP + Phosphoribosylformylglycineamidine + L-Glutamate + Hydrogen ion + Phosphate Adenosine triphosphate + L-Glutamine + Water + Uridine triphosphate > ADP + Cytidine triphosphate + L-Glutamate +2 Hydrogen ion + Phosphate Fructose 6-phosphate + L-Glutamine <> Glucosamine 6-phosphate + L-Glutamate 2 L-Glutamate + NAD <> L-Glutamine + alpha-Ketoglutarate + NADH + Hydrogen ion 2 L-Glutamate + NADP <> L-Glutamine + alpha-Ketoglutarate + NADPH + Hydrogen ion Adenosine triphosphate + L-Glutamate + Ammonia <> ADP + Phosphate + L-Glutamine L-Glutamine + Water <> L-Glutamate + Ammonia Adenosine triphosphate + Uridine triphosphate + L-Glutamine + Water <> ADP + Phosphate + Cytidine triphosphate + L-Glutamate 2 Adenosine triphosphate + L-Glutamine + Hydrogen carbonate + Water <>2 ADP + Phosphate + L-Glutamate + Carbamoylphosphate Adenosine triphosphate + L-Aspartic acid + L-Glutamine + Water <> Adenosine monophosphate + Pyrophosphate + L-Asparagine + L-Glutamate Chorismate + L-Glutamine <> 2-Aminobenzoic acid + Pyruvic acid + L-Glutamate More...Adenosine triphosphate + Xanthylic acid + L-Glutamine + Water <> Adenosine monophosphate + Pyrophosphate + Guanosine monophosphate + L-Glutamate Adenosine triphosphate + L-Glutamine + tRNA(Gln) + tRNA(Gln) <> Adenosine monophosphate + Pyrophosphate + Glutaminyl-tRNA + Glutaminyl-tRNA Adenosine triphosphate + 5'-Phosphoribosyl-N-formylglycineamide + L-Glutamine + Water <> ADP + Phosphate + Phosphoribosylformylglycineamidine + L-Glutamate Phosphoribulosylformimino-AICAR-P + L-Glutamine <> D-Erythro-imidazole-glycerol-phosphate + AICAR + L-Glutamate 6-Thioxanthine 5'-monophosphate + Adenosine triphosphate + L-Glutamine + Water <> 6-Thioguanosine monophosphate + Adenosine monophosphate + Pyrophosphate + L-Glutamate -->-->L-Glutamate + NADP < Hydrogen ion + L-Glutamine + Oxoglutaric acid + NADPH Phosphoribulosylformimino-AICAR-P + L-Glutamine > Hydrogen ion + L-Glutamate + D-Erythro-imidazole-glycerol-phosphate + AICAR L-Glutamine + Water > Hydrogen ion + L-Glutamate + Ammonia Adenosine triphosphate + Nicotinic acid adenine dinucleotide + L-Glutamine + Water > Hydrogen ion + Adenosine monophosphate + Pyrophosphate + NAD + L-Glutamate 5-Phosphoribosylamine + Pyrophosphate + L-Glutamate < Phosphoribosyl pyrophosphate + L-Glutamine + Water ala-gln + Water > L-Alanine + L-Glutamine gly-gln + Water > Glycine + L-Glutamine 2 Adenosine triphosphate + L-Glutamine + Carbonic acid + Water >2 ADP + Inorganic phosphate + L-Glutamate + Carbamoylphosphate Adenosine triphosphate + L-Glutamate + Ammonia > ADP + Inorganic phosphate + L-Glutamine 2 L-Glutamate + NADP > L-Glutamine + Oxoglutaric acid + NADPH Adenosine triphosphate + Uridine triphosphate + L-Glutamine + Water + Ammonia <> ADP + Phosphate + Cytidine triphosphate + L-Glutamate 2 Adenosine triphosphate + L-Glutamine + Hydrogen carbonate + Water + Ammonia + Carbamic acid + Carboxyphosphate <>2 ADP + Phosphate + L-Glutamate + Carbamoylphosphate 2 L-Glutamate + NADP + Ammonia + Water <> L-Glutamine + alpha-Ketoglutarate + NADPH + Hydrogen ion Adenosine triphosphate + L-Aspartic acid + L-Glutamine + Water + Ammonia <> Adenosine monophosphate + Pyrophosphate + L-Asparagine + L-Glutamate Adenosine triphosphate + Xanthylic acid + L-Glutamine + Water + Ammonia <> Adenosine monophosphate + Pyrophosphate + Guanosine monophosphate + L-Glutamate Ammonia + L-Glutamic acid + Adenosine triphosphate + Oxoglutaric acid + L-Glutamate <> Phosphate + L-Glutamine + Adenosine diphosphate + ADP L-Glutamic acid + Adenosine triphosphate + Ammonium + L-Glutamate > L-Glutamine + Hydrogen ion + Adenosine diphosphate + Phosphate + ADP 2 L-Glutamic acid + NADP + 2 L-Glutamate > L-Glutamine + NADPH + Hydrogen ion + Oxoglutaric acid + NADPH L-Glutamine + Water > L-Glutamic acid + Ammonia + L-Glutamate Adenosine triphosphate + L-Aspartic acid + L-Glutamine + Water + L-Aspartic acid > Adenosine monophosphate + Pyrophosphate + L-Asparagine + L-Glutamic acid + L-Asparagine + L-Glutamate Hydrogen carbonate + Water + L-Glutamine + 2 Adenosine triphosphate >2 Adenosine diphosphate + Phosphate + L-Glutamic acid +2 Hydrogen ion + Carbamoylphosphate +2 ADP + L-Glutamate L-Glutamine + Adenosine triphosphate + Hydrogen ion + tRNA(Gln) > Adenosine diphosphate + Pyrophosphate + L-Glutaminyl-tRNA(Gln) + ADP Phosphoribulosylformimino-AICAR-P + L-Glutamine > L-Glutamic acid + Hydrogen ion + 5-Amino-4-imidazolecarboxyamide + D-Erythro-imidazole-glycerol-phosphate + L-Glutamate L-Aspartic acid + Water + Adenosine triphosphate + L-Glutamine + L-Aspartic acid > L-Asparagine + Hydrogen ion + Adenosine monophosphate + L-Glutamic acid + Pyrophosphate + L-Asparagine + L-Glutamate Chorismate + L-Glutamine > L-Glutamic acid + Pyruvic acid + Hydrogen ion + 2-Aminobenzoic acid + L-Glutamate Nicotinic acid adenine dinucleotide + Water + L-Glutamine + Adenosine triphosphate > Hydrogen ion + Adenosine monophosphate + Pyrophosphate + L-Glutamic acid + NAD + L-Glutamate D-tagatofuranose 6-phosphate + L-Glutamine <> Glucosamine 6-phosphate + L-Glutamic acid + L-Glutamate Fructose 6-phosphate + L-Glutamine + Fructose 6-phosphate > L-Glutamic acid + Glucosamine 6-phosphate + L-Glutamate Chorismate + L-Glutamine > L-Glutamic acid + 4-amino-4-deoxychorismate + L-Glutamate + 4-Amino-4-deoxychorismate Phosphoribosyl pyrophosphate + Water + L-Glutamine > 5-Phosphoribosylamine + L-Glutamic acid + Pyrophosphate + 5-Phosphoribosylamine + L-Glutamate 5'-Phosphoribosyl-N-formylglycinamide + Water + L-Glutamine + Adenosine triphosphate + 5'-Phosphoribosyl-N-formylglycineamide > 2-(Formamido)-N1-(5-phospho-D-ribosyl)acetamidine + L-Glutamic acid + Phosphate + Adenosine diphosphate + Hydrogen ion + L-Glutamate + ADP Xanthylic acid + Adenosine triphosphate + L-Glutamine + Water > Adenosine monophosphate + Pyrophosphate + L-Glutamic acid +2 Hydrogen ion + Guanosine monophosphate + L-Glutamate Uridine triphosphate + L-Glutamine + Water + Adenosine triphosphate + Uridine triphosphate > Adenosine diphosphate + Hydrogen ion + Phosphate + L-Glutamic acid + Cytidine triphosphate + ADP + L-Glutamate L-Glutamine + Adenosine triphosphate + Water > Adenosine diphosphate + Phosphate + Hydrogen ion + L-Glutamine + ADP L-Glutamine + Adenosine triphosphate + Water > Adenosine diphosphate + Phosphate + Hydrogen ion + L-Glutamine + ADP | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Pathways: |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Spectra | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Spectra: | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| References | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| References: |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Synthesis Reference: | Jiao, Qingcai; Qian, Shaosong; Chen, Ran; Wu, Xiaoyan. Synthesis of L-glutamine. Faming Zhuanli Shenqing Gongkai Shuomingshu (2005), 7 pp. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Material Safety Data Sheet (MSDS) | Download (PDF) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Links | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| External Links: |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
Enzymes
- General function:
- Involved in asparaginase activity
- Specific function:
- L-asparagine + H(2)O = L-aspartate + NH(3)
- Gene Name:
- ansB
- Locus Tag:
- PA1337
- Molecular weight:
- 38.6 kDa
Reactions
| L-asparagine + H(2)O = L-aspartate + NH(3). |
- General function:
- Involved in biosynthetic process
- Specific function:
- Chorismate + L-glutamine = anthranilate + pyruvate + L-glutamate
- Gene Name:
- trpE
- Locus Tag:
- PA0609
- Molecular weight:
- 54.6 kDa
Reactions
| Chorismate + L-glutamine = anthranilate + pyruvate + L-glutamate. |
- General function:
- Involved in metabolic process
- Specific function:
- Catalyzes the biosynthesis of 4-amino-4-deoxychorismate (ADC) from chorismate and glutamine
- Gene Name:
- pabA
- Locus Tag:
- PA0649
- Molecular weight:
- 22 kDa
Reactions
| Chorismate + L-glutamine = 4-amino-4-deoxychorismate + L-glutamate. |
- General function:
- Involved in anthranilate phosphoribosyltransferase activity
- Specific function:
- Chorismate + L-glutamine = anthranilate + pyruvate + L-glutamate
- Gene Name:
- trpD
- Locus Tag:
- PA0650
- Molecular weight:
- 37.4 kDa
Reactions
| Chorismate + L-glutamine = anthranilate + pyruvate + L-glutamate. |
| N-(5-phospho-D-ribosyl)-anthranilate + diphosphate = anthranilate + 5-phospho-alpha-D-ribose 1-diphosphate. |
- General function:
- Involved in nucleotide binding
- Specific function:
- ATP + L-glutamine + tRNA(Gln) = AMP + diphosphate + L-glutaminyl-tRNA(Gln)
- Gene Name:
- glnS
- Locus Tag:
- PA1794
- Molecular weight:
- 62.9 kDa
Reactions
| ATP + L-glutamine + tRNA(Gln) = AMP + diphosphate + L-glutaminyl-tRNA(Gln). |
- General function:
- Involved in ATP binding
- Specific function:
- 2 ATP + L-glutamine + HCO(3)(-) + H(2)O = 2 ADP + phosphate + L-glutamate + carbamoyl phosphate
- Gene Name:
- carB
- Locus Tag:
- PA4756
- Molecular weight:
- 117.3 kDa
Reactions
| 2 ATP + L-glutamine + HCO(3)(-) + H(2)O = 2 ADP + phosphate + L-glutamate + carbamoyl phosphate. |
- General function:
- Involved in GMP synthase (glutamine-hydrolyzing) activity
- Specific function:
- Catalyzes the synthesis of GMP from XMP
- Gene Name:
- guaA
- Locus Tag:
- PA3769
- Molecular weight:
- 58 kDa
Reactions
| ATP + xanthosine 5'-phosphate + L-glutamine + H(2)O = AMP + diphosphate + GMP + L-glutamate. |
- General function:
- Involved in biosynthetic process
- Specific function:
- Catalyzes the biosynthesis of 4-amino-4-deoxychorismate (ADC) from chorismate and glutamine
- Gene Name:
- pabB
- Locus Tag:
- PA1758
- Molecular weight:
- 49.6 kDa
Reactions
| Chorismate + L-glutamine = 4-amino-4-deoxychorismate + L-glutamate. |
- General function:
- Involved in catalytic activity
- Specific function:
- 2 L-glutamate + NADP(+) = L-glutamine + 2- oxoglutarate + NADPH
- Gene Name:
- gltB
- Locus Tag:
- PA5036
- Molecular weight:
- 161.6 kDa
Reactions
| 2 L-glutamate + NADP(+) = L-glutamine + 2-oxoglutarate + NADPH. |
- General function:
- Involved in iron-sulfur cluster binding
- Specific function:
- 2 L-glutamate + NADP(+) = L-glutamine + 2- oxoglutarate + NADPH
- Gene Name:
- gltD
- Locus Tag:
- PA5035
- Molecular weight:
- 52.6 kDa
Reactions
| 2 L-glutamate + NADP(+) = L-glutamine + 2-oxoglutarate + NADPH. |
- General function:
- Involved in glutamine catabolic process
- Specific function:
- 2 ATP + L-glutamine + HCO(3)(-) + H(2)O = 2 ADP + phosphate + L-glutamate + carbamoyl phosphate
- Gene Name:
- carA
- Locus Tag:
- PA4758
- Molecular weight:
- 40.8 kDa
Reactions
| 2 ATP + L-glutamine + HCO(3)(-) + H(2)O = 2 ADP + phosphate + L-glutamate + carbamoyl phosphate. |
- General function:
- Involved in glutaminase activity
- Specific function:
- L-glutamine + H(2)O = L-glutamate + NH(3)
- Gene Name:
- glsA2
- Locus Tag:
- PA1638
- Molecular weight:
- 33 kDa
Reactions
| L-glutamine + H(2)O = L-glutamate + NH(3). |
- General function:
- Involved in CTP synthase activity
- Specific function:
- Catalyzes the ATP-dependent amination of UTP to CTP with either L-glutamine or ammonia as the source of nitrogen
- Gene Name:
- pyrG
- Locus Tag:
- PA3637
- Molecular weight:
- 59.6 kDa
Reactions
| ATP + UTP + NH(3) = ADP + phosphate + CTP. |
- General function:
- Involved in glutamate-ammonia ligase activity
- Specific function:
- ATP + L-glutamate + NH(3) = ADP + phosphate + L-glutamine
- Gene Name:
- glnA
- Locus Tag:
- PA5119
- Molecular weight:
- 51.9 kDa
Reactions
| ATP + L-glutamate + NH(3) = ADP + phosphate + L-glutamine. |
- General function:
- Involved in amidophosphoribosyltransferase activity
- Specific function:
- 5-phospho-beta-D-ribosylamine + diphosphate + L-glutamate = L-glutamine + 5-phospho-alpha-D-ribose 1-diphosphate + H(2)O
- Gene Name:
- purF
- Locus Tag:
- PA3108
- Molecular weight:
- 55.4 kDa
Reactions
| 5-phospho-beta-D-ribosylamine + diphosphate + L-glutamate = L-glutamine + 5-phospho-alpha-D-ribose 1-diphosphate + H(2)O. |
- General function:
- Involved in catalytic activity
- Specific function:
- ATP + N(2)-formyl-N(1)-(5-phospho-D- ribosyl)glycinamide + L-glutamine + H(2)O = ADP + phosphate + 2- (formamido)-N(1)-(5-phospho-D-ribosyl)acetamidine + L-glutamate
- Gene Name:
- purL
- Locus Tag:
- PA3763
- Molecular weight:
- 140.6 kDa
Reactions
| ATP + N(2)-formyl-N(1)-(5-phospho-D-ribosyl)glycinamide + L-glutamine + H(2)O = ADP + phosphate + 2-(formamido)-N(1)-(5-phospho-D-ribosyl)acetamidine + L-glutamate. |
- General function:
- Involved in metabolic process
- Specific function:
- Catalyzes the first step in hexosamine metabolism, converting fructose-6P into glucosamine-6P using glutamine as a nitrogen source
- Gene Name:
- glmS
- Locus Tag:
- PA5549
- Molecular weight:
- 66.3 kDa
Reactions
| L-glutamine + D-fructose 6-phosphate = L-glutamate + D-glucosamine 6-phosphate. |
- General function:
- Involved in NAD+ synthase (glutamine-hydrolyzing) activity
- Specific function:
- Catalyzes a key step in NAD biosynthesis, transforming deamido-NAD into NAD by a two-step reaction
- Gene Name:
- nadE
- Locus Tag:
- PA4920
- Molecular weight:
- 29.7 kDa
Reactions
| ATP + deamido-NAD(+) + NH(3) = AMP + diphosphate + NAD(+). |
Transporters
- General function:
- Involved in nucleotide binding
- Specific function:
- Probably part of a binding-protein-dependent transport system yecCS for an amino acid. Probably responsible for energy coupling to the transport system
- Gene Name:
- yecC
- Locus Tag:
- PA5152
- Molecular weight:
- 28.4 kDa