Inosinic acid (PAMDB000601)
| Record Information | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Version | 1.0 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Update Date | 1/22/2018 12:54:54 PM | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Metabolite ID | PAMDB000601 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Identification | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
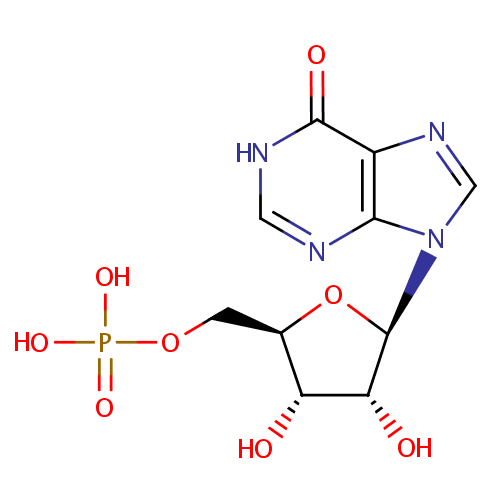
| Name: | Inosinic acid | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Description: | Inosinic acid is a purine nucleotide which has hypoxanthine as the base and one phosphate group esterified to the sugar moiety. It is formed by the deamination of AMP and when hydrolysed produces inosine. Inosinic acid is the ribonucleotide of hypoxanthine and is the first compound formed during the synthesis of purine. (Wikipedia) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Structure | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Synonyms: |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Chemical Formula: | C10H13N4O8P | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Average Molecular Weight: | 348.206 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Monoisotopic Molecular Weight: | 348.047099924 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| InChI Key: | GRSZFWQUAKGDAV-KQYNXXCUSA-N | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| InChI: | InChI=1S/C10H13N4O8P/c15-6-4(1-21-23(18,19)20)22-10(7(6)16)14-3-13-5-8(14)11-2-12-9(5)17/h2-4,6-7,10,15-16H,1H2,(H,11,12,17)(H2,18,19,20)/t4-,6-,7-,10-/m1/s1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| CAS number: | 131-99-7 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| IUPAC Name: | {[(2R,3S,4R,5R)-3,4-dihydroxy-5-(6-oxo-6,9-dihydro-1H-purin-9-yl)oxolan-2-yl]methoxy}phosphonic acid | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Traditional IUPAC Name: | inosine-5'-monophosphate | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| SMILES: | O[C@@H]1[C@@H](COP(O)(O)=O)O[C@H]([C@@H]1O)N1C=NC2=C1N=CNC2=O | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Chemical Taxonomy | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Taxonomy Description | This compound belongs to the class of organic compounds known as purine ribonucleoside monophosphates. These are nucleotides consisting of a purine base linked to a ribose to which one monophosphate group is attached. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Kingdom | Organic compounds | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Super Class | Nucleosides, nucleotides, and analogues | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Class | Purine nucleotides | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Sub Class | Purine ribonucleotides | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Direct Parent | Purine ribonucleoside monophosphates | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Alternative Parents |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Substituents |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Molecular Framework | Aromatic heteropolycyclic compounds | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| External Descriptors |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Physical Properties | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| State: | Solid | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Charge: | -2 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Melting point: | Not Available | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Experimental Properties: |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Predicted Properties |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Biological Properties | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Cellular Locations: | Cytoplasm | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Reactions: | Hypoxanthine + Phosphoribosyl pyrophosphate <> Inosinic acid + Pyrophosphate Water + Inosinic acid > Inosine + Phosphate Guanosine monophosphate + 2 Hydrogen ion + NADPH > Inosinic acid + NADP + Ammonium Adenosine triphosphate + Inosine <> ADP + Hydrogen ion + Inosinic acid Water + Inosinic acid + NAD <> Hydrogen ion + NADH + Xanthylic acid Water + Inosine triphosphate > Hydrogen ion + Inosinic acid + Pyrophosphate Water + Inosinic acid <> Phosphoribosyl formamidocarboxamide L-Aspartic acid + Guanosine triphosphate + Inosinic acid <> Adenylsuccinic acid + Guanosine diphosphate +2 Hydrogen ion + Phosphate Inosine triphosphate + Water <> Inosinic acid + Pyrophosphate Adenosine triphosphate + Inosine <> ADP + Inosinic acid Inosinic acid + Ammonia + NADP <> Guanosine monophosphate + NADPH + Hydrogen ion Guanosine triphosphate + Inosinic acid + L-Aspartic acid <> Guanosine diphosphate + Phosphate + Adenylsuccinic acid L-Aspartic acid + Inosinic acid + Guanosine triphosphate > Hydrogen ion + adenylo-succinate + Phosphate + Guanosine diphosphate Inosinic acid + Ammonia + NADP > Guanosine monophosphate + NADPH More...Inosinic acid + NAD + Water > Xanthylic acid + NADH Guanosine triphosphate + Inosinic acid + L-Aspartic acid + L-Aspartic acid > Guanosine diphosphate + Phosphate + N(6)-(1,2-dicarboxyethyl)AMP Inosinic acid + L-Aspartic acid + Guanosine triphosphate + L-Aspartic acid > Guanosine diphosphate + Phosphate +2 Hydrogen ion + N(6)-(1,2-dicarboxyethyl)AMP + Adenylsuccinic acid FAICAR <> Inosinic acid + Water | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Pathways: | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Spectra | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Spectra: | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| References | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| References: |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Synthesis Reference: | Park, Yeong Hun; Cho, Gwang Myeong; Baek, Min Ji; Hong, Guk Gi; Lee, Jin Nam. Method for preparing 5'-inosinic acid by using microbe capable of over-expressing purC gene. Repub. Korea (2007), 7pp. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Material Safety Data Sheet (MSDS) | Download (PDF) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Links | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| External Links: |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
Enzymes
- General function:
- Involved in adenylosuccinate synthase activity
- Specific function:
- Plays an important role in the de novo pathway of purine nucleotide biosynthesis. Catalyzes the first commited step in the biosynthesis of AMP from IMP
- Gene Name:
- purA
- Locus Tag:
- PA4938
- Molecular weight:
- 46.8 kDa
Reactions
| GTP + IMP + L-aspartate = GDP + phosphate + N(6)-(1,2-dicarboxyethyl)-AMP. |
- General function:
- Involved in hydrolase activity
- Specific function:
- Nucleotidase with a broad substrate specificity as it can dephosphorylate various ribo- and deoxyribonucleoside 5'- monophosphates and ribonucleoside 3'-monophosphates with highest affinity to 3'-AMP. Also hydrolyzes polyphosphate (exopolyphosphatase activity) with the preference for short-chain- length substrates (P20-25). Might be involved in the regulation of dNTP and NTP pools, and in the turnover of 3'-mononucleotides produced by numerous intracellular RNases (T1, T2, and F) during the degradation of various RNAs. Also plays a significant physiological role in stress-response and is required for the survival of Pseudomonas aeruginosa in stationary growth phase
- Gene Name:
- surE
- Locus Tag:
- PA3625
- Molecular weight:
- 26.4 kDa
Reactions
| A 5'-ribonucleotide + H(2)O = a ribonucleoside + phosphate. |
| A 3'-ribonucleotide + H(2)O = a ribonucleoside + phosphate. |
| (Polyphosphate)(n) + H(2)O = (polyphosphate)(n-1) + phosphate. |
- General function:
- Involved in catalytic activity
- Specific function:
- Inosine 5'-phosphate + NAD(+) + H(2)O = xanthosine 5'-phosphate + NADH
- Gene Name:
- guaB
- Locus Tag:
- PA3770
- Molecular weight:
- 51.7 kDa
Reactions
| Inosine 5'-phosphate + NAD(+) + H(2)O = xanthosine 5'-phosphate + NADH. |
- General function:
- Involved in acid phosphatase activity
- Specific function:
- Dephosphorylates several organic phosphomonoesters and catalyzes the transfer of low-energy phosphate groups from phosphomonoesters to hydroxyl groups of various organic compounds. Preferentially acts on aryl phosphoesters. Might function as a broad-spectrum dephosphorylating enzyme able to scavenge both 3'- and 5'-nucleotides and also additional organic phosphomonoesters
- Gene Name:
- aphA
- Locus Tag:
- PA1409
- Molecular weight:
- 38 kDa
Reactions
| A phosphate monoester + H(2)O = an alcohol + phosphate. |
- General function:
- Involved in nucleoside-triphosphate diphosphatase activity
- Specific function:
- Specific function unknown
- Gene Name:
- mazG
- Locus Tag:
- PA0935
- Molecular weight:
- 31.2 kDa
Reactions
| ATP + H(2)O = AMP + diphosphate. |
- General function:
- Involved in IMP cyclohydrolase activity
- Specific function:
- 10-formyltetrahydrofolate + 5-amino-1-(5- phospho-D-ribosyl)imidazole-4-carboxamide = tetrahydrofolate + 5- formamido-1-(5-phospho-D-ribosyl)imidazole-4-carboxamide
- Gene Name:
- purH
- Locus Tag:
- PA4854
- Molecular weight:
- 57.7 kDa
Reactions
| 10-formyltetrahydrofolate + 5-amino-1-(5-phospho-D-ribosyl)imidazole-4-carboxamide = tetrahydrofolate + 5-formamido-1-(5-phospho-D-ribosyl)imidazole-4-carboxamide. |
| IMP + H(2)O = 5-formamido-1-(5-phospho-D-ribosyl)imidazole-4-carboxamide. |
- General function:
- Involved in hydrolase activity
- Specific function:
- Hydrolyzes O6 atom-containing purine bases deoxyinosine triphosphate (dITP) and xanthosine triphosphate (XTP) as well as 2'-deoxy-N-6-hydroxylaminopurine triposphate (dHAPTP) to nucleotide monophosphate and pyrophosphate. Probably excludes non- standard purines from DNA precursor pool, preventing thus incorporation into DNA and avoiding chromosomal lesions
- Gene Name:
- rdgB
- Locus Tag:
- PA0387
- Molecular weight:
- 21.2 kDa
Reactions
| A nucleoside triphosphate + H(2)O = a nucleotide + diphosphate. |