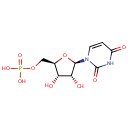
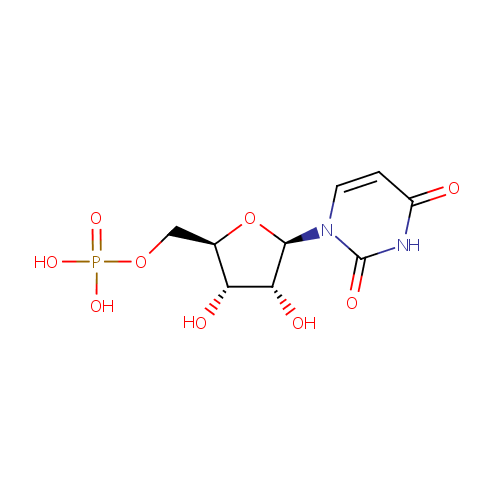
Uridine 5'-monophosphate (PAMDB000118)
Enzymes
- General function:
- Involved in catalytic activity
- Specific function:
- Catalyzes the decarboxylation of orotidine 5'- monophosphate (OMP) to uridine 5'-monophosphate (UMP)
- Gene Name:
- pyrF
- Locus Tag:
- PA2876
- Molecular weight:
- 24.4 kDa
Reactions
| Orotidine 5'-phosphate = UMP + CO(2). |
- General function:
- Involved in cytidylate kinase activity
- Specific function:
- ATP, dATP, and GTP are equally effective as phosphate donors. CMP and dCMP are the best phosphate acceptors
- Gene Name:
- cmk
- Locus Tag:
- PA3163
- Molecular weight:
- 24.6 kDa
Reactions
| ATP + (d)CMP = ADP + (d)CDP. |
- General function:
- Involved in phospho-N-acetylmuramoyl-pentapeptide-transferase activity
- Specific function:
- First step of the lipid cycle reactions in the biosynthesis of the cell wall peptidoglycan
- Gene Name:
- mraY
- Locus Tag:
- PA4415
- Molecular weight:
- 39.7 kDa
Reactions
| UDP-Mur2Ac(oyl-L-Ala-gamma-D-Glu-L-Lys-D-Ala-D-Ala) + undecaprenyl phosphate = UMP + Mur2Ac(oyl-L-Ala-gamma-D-Glu-L-Lys-D-Ala-D-Ala)-diphosphoundecaprenol. |
- General function:
- Involved in cellular amino acid biosynthetic process
- Specific function:
- Catalyzes the reversible phosphorylation of UMP to UDP, with ATP as the most efficient phosphate donor
- Gene Name:
- pyrH
- Locus Tag:
- PA3654
- Molecular weight:
- 26.3 kDa
Reactions
| ATP + UMP = ADP + UDP. |
- General function:
- Involved in hydrolase activity
- Specific function:
- Nucleotidase with a broad substrate specificity as it can dephosphorylate various ribo- and deoxyribonucleoside 5'- monophosphates and ribonucleoside 3'-monophosphates with highest affinity to 3'-AMP. Also hydrolyzes polyphosphate (exopolyphosphatase activity) with the preference for short-chain- length substrates (P20-25). Might be involved in the regulation of dNTP and NTP pools, and in the turnover of 3'-mononucleotides produced by numerous intracellular RNases (T1, T2, and F) during the degradation of various RNAs. Also plays a significant physiological role in stress-response and is required for the survival of Pseudomonas aeruginosa in stationary growth phase
- Gene Name:
- surE
- Locus Tag:
- PA3625
- Molecular weight:
- 26.4 kDa
Reactions
| A 5'-ribonucleotide + H(2)O = a ribonucleoside + phosphate. |
| A 3'-ribonucleotide + H(2)O = a ribonucleoside + phosphate. |
| (Polyphosphate)(n) + H(2)O = (polyphosphate)(n-1) + phosphate. |
- General function:
- Involved in nucleoside metabolic process
- Specific function:
- Catalyzes the conversion of uracil and 5-phospho-alpha- D-ribose 1-diphosphate (PRPP) to UMP and diphosphate
- Gene Name:
- upp
- Locus Tag:
- PA4646
- Molecular weight:
- 22.9 kDa
Reactions
| UMP + diphosphate = uracil + 5-phospho-alpha-D-ribose 1-diphosphate. |
- General function:
- Involved in acid phosphatase activity
- Specific function:
- Dephosphorylates several organic phosphomonoesters and catalyzes the transfer of low-energy phosphate groups from phosphomonoesters to hydroxyl groups of various organic compounds. Preferentially acts on aryl phosphoesters. Might function as a broad-spectrum dephosphorylating enzyme able to scavenge both 3'- and 5'-nucleotides and also additional organic phosphomonoesters
- Gene Name:
- aphA
- Locus Tag:
- PA1409
- Molecular weight:
- 38 kDa
Reactions
| A phosphate monoester + H(2)O = an alcohol + phosphate. |
- General function:
- Involved in nucleoside-triphosphate diphosphatase activity
- Specific function:
- Specific function unknown
- Gene Name:
- mazG
- Locus Tag:
- PA0935
- Molecular weight:
- 31.2 kDa
Reactions
| ATP + H(2)O = AMP + diphosphate. |
- General function:
- Involved in hydrolase activity
- Specific function:
- Hydrolyzes O6 atom-containing purine bases deoxyinosine triphosphate (dITP) and xanthosine triphosphate (XTP) as well as 2'-deoxy-N-6-hydroxylaminopurine triposphate (dHAPTP) to nucleotide monophosphate and pyrophosphate. Probably excludes non- standard purines from DNA precursor pool, preventing thus incorporation into DNA and avoiding chromosomal lesions
- Gene Name:
- rdgB
- Locus Tag:
- PA0387
- Molecular weight:
- 21.2 kDa
Reactions
| A nucleoside triphosphate + H(2)O = a nucleotide + diphosphate. |