L-Tyrosine (PAMDB000060)
| Record Information | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Version | 1.0 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Update Date | 1/22/2018 12:54:54 PM | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Metabolite ID | PAMDB000060 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Identification | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
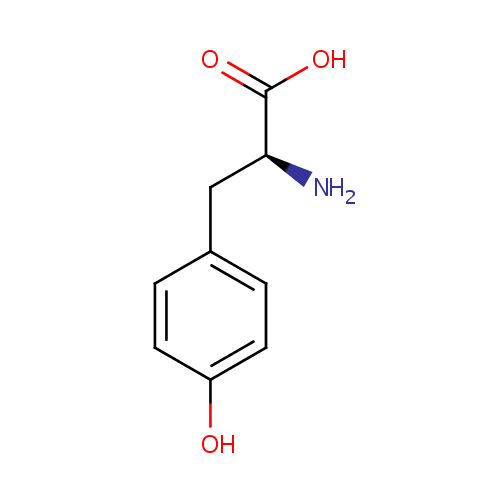
| Name: | L-Tyrosine | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Description: | Tyrosine (Tyr, Y) or 4-hydroxyphenylalanine is a non-essential amino acid with a polar side group. Its codons are UAC and UAU. Aside from being a proteogenic amino acid, tyrosine has a special role by virtue of the phenol functionality. It occurs in proteins that are part of signal transduction processes. It functions as a receiver of phosphate groups that are transferred by way of protein kinases (so-called receptor tyrosine kinases). Phosphorylation of the hydroxyl group changes the activity of the target protein. (Wikipedia) L-Tyrosine is the enantiomer of tyrosine (the other being D-tyrosine) that is used in building proteins. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Structure | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Synonyms: |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Chemical Formula: | C9H11NO3 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Average Molecular Weight: | 181.1885 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Monoisotopic Molecular Weight: | 181.073893223 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| InChI Key: | OUYCCCASQSFEME-QMMMGPOBSA-N | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| InChI: | InChI=1S/C9H11NO3/c10-8(9(12)13)5-6-1-3-7(11)4-2-6/h1-4,8,11H,5,10H2,(H,12,13)/t8-/m0/s1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| CAS number: | 60-18-4 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| IUPAC Name: | (2S)-2-amino-3-(4-hydroxyphenyl)propanoic acid | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Traditional IUPAC Name: | L-tyrosine | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| SMILES: | N[C@@H](CC1=CC=C(O)C=C1)C(O)=O | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Chemical Taxonomy | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Taxonomy Description | This compound belongs to the class of organic compounds known as phenylpropanoic acids. These are compounds with a structure containing a benzene ring conjugated to a propanoic acid. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Kingdom | Organic compounds | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Super Class | Phenylpropanoids and polyketides | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Class | Phenylpropanoic acids | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Sub Class | Not Available | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Direct Parent | Phenylpropanoic acids | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Alternative Parents | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Substituents |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Molecular Framework | Aromatic homomonocyclic compounds | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| External Descriptors |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Physical Properties | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| State: | Solid | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Charge: | 0 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Melting point: | 343 °C | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Experimental Properties: |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Predicted Properties |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Biological Properties | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Cellular Locations: | Cytoplasm | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Reactions: | alpha-Ketoglutarate + L-Tyrosine <> 4-Hydroxyphenylpyruvic acid + L-Glutamate Adenosine triphosphate + tRNA(Tyr) + L-Tyrosine + tRNA(Tyr) <> Adenosine monophosphate + Pyrophosphate + L-Tyrosyl-tRNA(Tyr) + L-Tyrosyl-tRNA(Tyr) S-Adenosylmethionine + NADPH + L-Tyrosine > p-Cresol + 5'-Deoxyadenosine + Dehydroglycine + Hydrogen ion + L-Methionine + NADP Water + Phosphotyrosine > Phosphate + L-Tyrosine Adenosine triphosphate + L-Tyrosine + tRNA(Tyr) <> Adenosine monophosphate + Pyrophosphate + L-Tyrosyl-tRNA(Tyr) C15815 + L-Tyrosine + Iminoglycine <> 4-Methyl-5-(2-phosphoethyl)-thiazole L-Tyrosine + S-Adenosylmethionine + a reduced electron acceptor > Dehydroglycine + p-Cresol + 5'-Deoxyadenosine + L-Methionine + an oxidized electron acceptor + Hydrogen ion L-Tyrosine + Oxoglutaric acid <> 4-Hydroxyphenylpyruvic acid + L-Glutamate L-Tyrosine + S-Adenosylmethionine + NADPH <> 2-iminoacetate + p-Cresol + 5'-Deoxyadenosine + L-Methionine + NADP + Hydrogen ion L-Tyrosine + Adenosine triphosphate + Hydrogen ion + tRNA(Tyr) + L-Tyrosine > Adenosine monophosphate + Pyrophosphate + L-tyrosyl-tRNA(Tyr) 4-hydroxyphenylpyruvate + L-Glutamic acid + L-Glutamate > Oxoglutaric acid + L-Tyrosine + L-Tyrosine L-Tyrosine + NADPH + S-adenosyl-L-methionine + L-Tyrosine + NADPH > Hydrogen ion + NADP + L-Methionine + 5'-Deoxyadenosine + p-Cresol + 2-iminoacetate | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Pathways: |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Spectra | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Spectra: | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| References | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| References: |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Synthesis Reference: | Enei, Hitoshi; Matsui, Hiroshi; Yamashita, Koichi; Okumura, Shinji; Yamada, Hideaki. Microbiological synthesis of L-tyrosine and 3,4-dihydroxyphenyl-L-alanine. I. Distribution of tyrosine phenol lyase in microorganisms. Agricultural and Biological Chemist | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Material Safety Data Sheet (MSDS) | Download (PDF) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Links | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| External Links: |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
Enzymes
- General function:
- Involved in transferase activity
- Specific function:
- An aromatic amino acid + 2-oxoglutarate = an aromatic oxo acid + L-glutamate
- Gene Name:
- tyrB
- Locus Tag:
- PA3139
- Molecular weight:
- 43.3 kDa
Reactions
| An aromatic amino acid + 2-oxoglutarate = an aromatic oxo acid + L-glutamate. |
- General function:
- Involved in acid phosphatase activity
- Specific function:
- Dephosphorylates several organic phosphomonoesters and catalyzes the transfer of low-energy phosphate groups from phosphomonoesters to hydroxyl groups of various organic compounds. Preferentially acts on aryl phosphoesters. Might function as a broad-spectrum dephosphorylating enzyme able to scavenge both 3'- and 5'-nucleotides and also additional organic phosphomonoesters
- Gene Name:
- aphA
- Locus Tag:
- PA1409
- Molecular weight:
- 38 kDa
Reactions
| A phosphate monoester + H(2)O = an alcohol + phosphate. |
- General function:
- Involved in nucleotide binding
- Specific function:
- Catalyzes the attachment of tyrosine to tRNA(Tyr) in a two-step reaction:tyrosine is first activated by ATP to form Tyr- AMP and then transferred to the acceptor end of tRNA(Tyr)
- Gene Name:
- tyrS
- Locus Tag:
- PA4138
- Molecular weight:
- 45.1 kDa
Reactions
| ATP + L-tyrosine + tRNA(Tyr) = AMP + diphosphate + L-tyrosyl-tRNA(Tyr). |
- General function:
- Involved in catalytic activity
- Specific function:
- Catalyzes the rearrangement of 1-deoxy-D-xylulose 5- phosphate (DXP) to produce the thiazole phosphate moiety of thiamine. Sulfur is provided by the thiocarboxylate moiety of the carrier protein ThiS. In vitro, sulfur can be provided by H(2)S
- Gene Name:
- thiG
- Locus Tag:
- PA0381
- Molecular weight:
- 28.2 kDa
Reactions
| 1-deoxy-D-xylulose 5-phosphate + 2-iminoacetate + thiocarboxy-adenylate-[sulfur-carrier protein ThiS] = 2-((2R,5Z)-2-carboxy-4-methylthiazol-5(2H)-ylidene)ethyl phosphate + [sulfur-carrier protein ThiS] + 2 H(2)O. |
Transporters
- General function:
- Involved in nucleotide binding
- Specific function:
- Probably part of a binding-protein-dependent transport system yecCS for an amino acid. Probably responsible for energy coupling to the transport system
- Gene Name:
- yecC
- Locus Tag:
- PA5152
- Molecular weight:
- 28.4 kDa
- General function:
- Involved in transport
- Specific function:
- Permease that is involved in the transport across the cytoplasmic membrane of the aromatic amino acids (phenylalanine, tyrosine, and tryptophan)
- Gene Name:
- aroP
- Locus Tag:
- PA3000
- Molecular weight:
- 51 kDa