FAD (PAMDB000308)
| Record Information | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Version | 1.0 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Update Date | 1/22/2018 12:54:54 PM | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Metabolite ID | PAMDB000308 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Identification | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
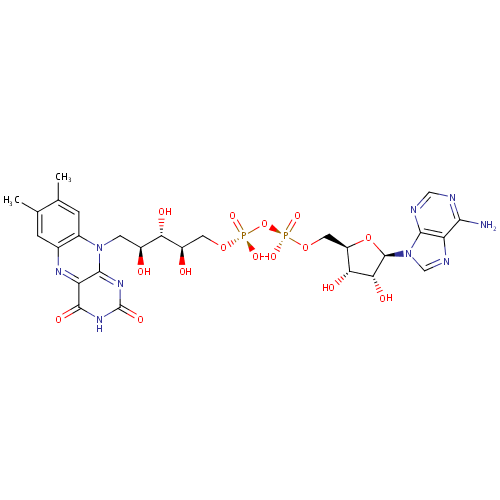
| Name: | FAD | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Description: | Flavine Adenine Dinucleotide (FAD) is a condensation product of riboflavin and adenosine diphosphate. FAD is a redox cofactor involved in several important reactions in metabolism. It can exist in two different redox states, (FAD and FADH2) which it converts between by accepting or donating electrons. It is a coenzyme of various aerobic dehydrogenases, e.g., D-amino acid oxidase and L-amino acid oxidase. (Lehninger, Principles of Biochemistry, 1982, p972) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Structure | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Synonyms: |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Chemical Formula: | C27H33N9O15P2 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Average Molecular Weight: | 785.5497 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Monoisotopic Molecular Weight: | 785.157134455 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| InChI Key: | VWWQXMAJTJZDQX-UYBVJOGSSA-N | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| InChI: | InChI=1S/C27H33N9O15P2/c1-10-3-12-13(4-11(10)2)35(24-18(32-12)25(42)34-27(43)33-24)5-14(37)19(39)15(38)6-48-52(44,45)51-53(46,47)49-7-16-20(40)21(41)26(50-16)36-9-31-17-22(28)29-8-30-23(17)36/h3-4,8-9,14-16,19-21,26,37-41H,5-7H2,1-2H3,(H,44,45)(H,46,47)(H2,28,29,30)(H,34,42,43)/t14-,15+,16+,19-,20+,21+,26+/m0/s1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| CAS number: | 146-14-5 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| IUPAC Name: | [({[(2R,3S,4R,5R)-5-(6-amino-9H-purin-9-yl)-3,4-dihydroxyoxolan-2-yl]methoxy}(hydroxy)phosphoryl)oxy]({[(2R,3S,4S)-5-{7,8-dimethyl-2,4-dioxo-2H,3H,4H,10H-benzo[g]pteridin-10-yl}-2,3,4-trihydroxypentyl]oxy})phosphinic acid | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Traditional IUPAC Name: | flavine-adenine dinucleotide | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| SMILES: | CC1=CC2=C(C=C1C)N(C[C@H](O)[C@H](O)[C@H](O)CO[P@](O)(=O)O[P@@](O)(=O)OC[C@H]1O[C@H]([C@H](O)[C@@H]1O)N1C=NC3=C1N=CN=C3N)C1=NC(=O)NC(=O)C1=N2 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Chemical Taxonomy | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Taxonomy Description | This compound belongs to the class of organic compounds known as flavin nucleotides. These are nucleotides containing a flavin moiety. Flavin is a compound that contains the tricyclic isoalloxazine ring system, which bears 2 oxo groups at the 2- and 4-positions. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Kingdom | Organic compounds | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Super Class | Nucleosides, nucleotides, and analogues | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Class | Flavin nucleotides | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Sub Class | Not Available | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Direct Parent | Flavin nucleotides | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Alternative Parents |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Substituents |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Molecular Framework | Aromatic heteropolycyclic compounds | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| External Descriptors |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Physical Properties | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| State: | Solid | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Charge: | -3 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Melting point: | Not Available | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Experimental Properties: |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Predicted Properties |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Biological Properties | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Cellular Locations: | Cytoplasm | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Reactions: | FADH2 + 2 Hydrogen ion + SufBCD with two bound [2Fe-2S] clusters > FAD + SufBCD with bound [4Fe-4S] cluster FAD + Hydrogen ion + NADPH > FADH2 + NADP Adenosine triphosphate + FADH2 + 2 Iron + Water + SufBCD scaffold complex + 2 SufSE with bound sulfur > ADP + FAD +7 Hydrogen ion + Phosphate + SufBCD with bound [2Fe-2S] cluster +2 SufSE sulfur acceptor complex Adenosine triphosphate + FADH2 + 2 Iron + Water + SufBCD with bound [2Fe-2S] cluster + 2 SufSE with bound sulfur > ADP + FAD +7 Hydrogen ion + Phosphate + SufBCD with two bound [2Fe-2S] clusters +2 SufSE sulfur acceptor complex FADH2 + 2 Iron + 2 IscS with bound sulfur + IscU scaffold protein > FAD +6 Hydrogen ion +2 IscS sulfur acceptor protein + IscU with bound [2Fe-2S] cluster FADH2 + 2 Iron + 2 IscS with bound sulfur + IscU with bound [2Fe-2S] cluster > FAD +6 Hydrogen ion +2 IscS sulfur acceptor protein + IscU with two bound [2Fe-2S] clusters Adenosine triphosphate + Flavin Mononucleotide + Hydrogen ion > FAD + Pyrophosphate Butyryl-CoA + FAD <> Crotonoyl-CoA + FADH2 FAD + Octanoyl-CoA <> FADH2 + (2E)-Octenoyl-CoA FAD + Palmityl-CoA <> FADH2 + (2E)-Hexadecenoyl-CoA FAD + Tetradecanoyl-CoA <> FADH2 + (2E)-Tetradecenoyl-CoA FAD + Hexanoyl-CoA <> FADH2 + trans-2-Hexenoyl-CoA FAD + Stearoyl-CoA <> FADH2 + Trans-Octadec-2-enoyl-CoA Lauroyl-CoA + FAD <> (2E)-Dodecenoyl-CoA + FADH2 Decanoyl-CoA (N-C10:0CoA) + FAD <> (2E)-Decenoyl-CoA + FADH2 FAD + L-Proline > L-D-1-Pyrroline-5-carboxylic acid + FADH2 + Hydrogen ion D-Alanine + FAD + Water > FADH2 + Ammonium + Pyruvic acid FADH2 + 2 Hydrogen ion + IscU with two bound [2Fe-2S] clusters > FAD + IscU with bound [4Fe-4S] cluster FADH2 + 2 Fe3+ > FAD +2 Iron +2 Hydrogen ion FAD + Hydrogen ion + NADH > FADH2 + NAD FADH2 + 2 Ferroxamine > FAD +2 Iron +2 ferroxamine minus Fe(3) +2 Hydrogen ion Adenosine triphosphate + Flavin Mononucleotide <> Pyrophosphate + FAD Succinic acid + FAD <> FADH2 + Fumaric acid Glycerol 3-phosphate + FAD <> Dihydroxyacetone phosphate + FADH2 Butanoyl-CoA + FAD <> FADH2 + Crotonoyl-CoA L-Malic acid + FAD <> FADH2 + Oxalacetic acid More...FAD + [Protein]-L-threonine <> [Protein]-FMN-L-Threonine + Adenosine monophosphate Succinic acid + Ubiquinone-10 + FAD <> Fumaric acid + QH2 + FADH2 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Pathways: | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Spectra | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Spectra: | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| References | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| References: |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Synthesis Reference: | Not Available | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Material Safety Data Sheet (MSDS) | Download (PDF) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Links | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| External Links: |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
Enzymes
- General function:
- Involved in electron carrier activity
- Specific function:
- Two distinct, membrane-bound, FAD-containing enzymes are responsible for the catalysis of fumarate and succinate interconversion; the fumarate reductase is used in anaerobic growth, and the succinate dehydrogenase is used in aerobic growth
- Gene Name:
- sdhB
- Locus Tag:
- PA1584
- Molecular weight:
- 26.2 kDa
Reactions
| Succinate + acceptor = fumarate + reduced acceptor. |
- General function:
- Involved in oxidoreductase activity
- Specific function:
- Oxidizes proline to glutamate for use as a carbon and nitrogen source and also function as a transcriptional repressor of the put operon
- Gene Name:
- putA
- Locus Tag:
- PA0782
- Molecular weight:
- 115.6 kDa
Reactions
| L-proline + acceptor = (S)-1-pyrroline-5-carboxylate + reduced acceptor. |
| (S)-1-pyrroline-5-carboxylate + NAD(P)(+) + 2 H(2)O = L-glutamate + NAD(P)H. |
- General function:
- Involved in metabolic process
- Specific function:
- Catalyzes the removal of elemental sulfur and selenium atoms from cysteine and selenocysteine to produce alanine. Functions as a sulfur delivery protein for NAD, biotin and Fe-S cluster synthesis. Transfers sulfur on 'Cys-456' of thiI in a transpersulfidation reaction. Transfers sulfur on 'Cys-19' of tusA in a transpersulfidation reaction. Functions also as a selenium delivery protein in the pathway for the biosynthesis of selenophosphate
- Gene Name:
- iscS
- Locus Tag:
- PA3814
- Molecular weight:
- 44.7 kDa
Reactions
| L-cysteine + acceptor = L-alanine + S-sulfanyl-acceptor. |
- General function:
- Involved in D-amino-acid dehydrogenase activity
- Specific function:
- Oxidative deamination of D-amino acids
- Gene Name:
- dadA
- Locus Tag:
- PA5304
- Molecular weight:
- 47.1 kDa
Reactions
| A D-amino acid + H(2)O + acceptor = a 2-oxo acid + NH(3) + reduced acceptor. |
- General function:
- Involved in electron carrier activity
- Specific function:
- Two distinct, membrane-bound, FAD-containing enzymes are responsible for the catalysis of fumarate and succinate interconversion; the fumarate reductase is used in anaerobic growth, and the succinate dehydrogenase is used in aerobic growth
- Gene Name:
- sdhA
- Locus Tag:
- PA1583
- Molecular weight:
- 63.5 kDa
Reactions
| Succinate + acceptor = fumarate + reduced acceptor. |
- General function:
- Involved in succinate dehydrogenase activity
- Specific function:
- Membrane-anchoring subunit of succinate dehydrogenase (SDH)
- Gene Name:
- sdhD
- Locus Tag:
- PA1582
- Molecular weight:
- 13.7 kDa
- General function:
- Involved in FMN adenylyltransferase activity
- Specific function:
- ATP + riboflavin = ADP + FMN
- Gene Name:
- ribF
- Locus Tag:
- PA4561
- Molecular weight:
- 34.3 kDa
Reactions
| ATP + riboflavin = ADP + FMN. |
| ATP + FMN = diphosphate + FAD. |
- General function:
- Involved in oxidoreductase activity
- Specific function:
- Conversion of glycerol 3-phosphate to dihydroxyacetone. Uses molecular oxygen or nitrate as electron acceptor
- Gene Name:
- glpD
- Locus Tag:
- PA3584
- Molecular weight:
- 57.1 kDa
Reactions
| sn-glycerol 3-phosphate + a quinone = glycerone phosphate + a quinol. |
- General function:
- Involved in sulfite reductase (NADPH) activity
- Specific function:
- Component of the sulfite reductase complex that catalyzes the 6-electron reduction of sulfite to sulfide. This is one of several activities required for the biosynthesis of L- cysteine from sulfate
- Gene Name:
- cysI
- Locus Tag:
- PA1838
- Molecular weight:
- 62.1 kDa
Reactions
| H(2)S + 3 NADP(+) + 3 H(2)O = sulfite + 3 NADPH. |
- General function:
- Involved in malate dehydrogenase (quinone) activity
- Specific function:
- (S)-malate + a quinone = oxaloacetate + reduced quinone
- Gene Name:
- mqo
- Locus Tag:
- PA3452
- Molecular weight:
- 57.2 kDa
Reactions
| (S)-malate + a quinone = oxaloacetate + reduced quinone. |
- General function:
- Involved in succinate dehydrogenase activity
- Specific function:
- Membrane-anchoring subunit of succinate dehydrogenase (SDH)
- Gene Name:
- sdhC
- Locus Tag:
- PA1581
- Molecular weight:
- 13.7 kDa
- General function:
- Involved in acyl-CoA dehydrogenase activity
- Specific function:
- Catalyzes the dehydrogenation of acyl-CoA
- Gene Name:
- fadE
- Locus Tag:
- PA2815
- Molecular weight:
- 88.8 kDa
Reactions
| An acyl-CoA + FAD = a dehydrogenated acyl-CoA + FADH(2). |
- General function:
- Involved in iron ion binding
- Specific function:
- May be involved in the formation or repair of [Fe-S] clusters present in iron-sulfur proteins (Potential)
- Gene Name:
- nifU
- Locus Tag:
- PA3813
- Molecular weight:
- 13.8 kDa