Glycine (PAMDB000046)
| Record Information | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Version | 1.0 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Update Date | 1/22/2018 12:54:54 PM | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Metabolite ID | PAMDB000046 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Identification | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
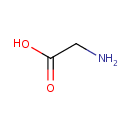
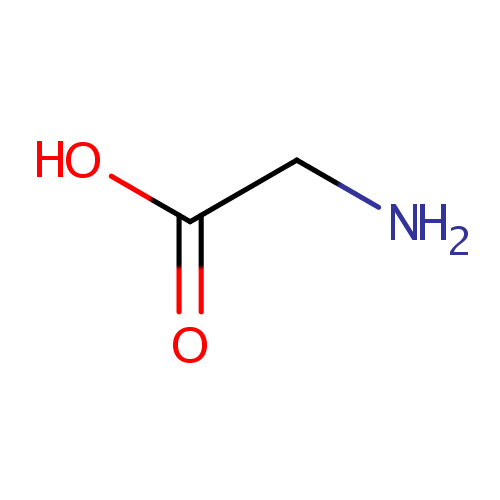
| Name: | Glycine | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Description: | Glycine is a simple amino acid. The glycine cleavage enzyme system comprises four proteins: P-, T-, H- and L-proteins (EC 1.4.4.2, EC 2.1.2.10 and EC 1.8.1.4 for P-, T- and L-proteins). The glycine cleavage system catalyses the oxidative conversion of glycine into carbon dioxide and ammonia, with the remaining one-carbon unit transferred to folate as methylenetetrahydrofolate. It is the main catabolic pathway for glycine and it also contributes to one-carbon metabolism. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Structure | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Synonyms: |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Chemical Formula: | C2H5NO2 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Average Molecular Weight: | 75.0666 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Monoisotopic Molecular Weight: | 75.032028409 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| InChI Key: | DHMQDGOQFOQNFH-UHFFFAOYSA-N | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| InChI: | InChI=1S/C2H5NO2/c3-1-2(4)5/h1,3H2,(H,4,5) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| CAS number: | 56-40-6 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| IUPAC Name: | 2-aminoacetic acid | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Traditional IUPAC Name: | glycine | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| SMILES: | NCC(O)=O | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Chemical Taxonomy | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Taxonomy Description | This compound belongs to the class of organic compounds known as alpha amino acids. These are amino acids in which the amino group is attached to the carbon atom immediately adjacent to the carboxylate group (alpha carbon). | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Kingdom | Organic compounds | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Super Class | Organic acids and derivatives | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Class | Carboxylic acids and derivatives | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Sub Class | Amino acids, peptides, and analogues | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Direct Parent | Alpha amino acids | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Alternative Parents | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Substituents |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Molecular Framework | Aliphatic acyclic compounds | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| External Descriptors |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Physical Properties | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| State: | Solid | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Charge: | 0 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Melting point: | 504 deg F | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Experimental Properties: |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Predicted Properties |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Biological Properties | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Cellular Locations: | Cytoplasm | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Reactions: | Cysteinylglycine + Water > L-Cysteine + Glycine L-Threonine <> Acetaldehyde + Glycine L-Allothreonine > Acetaldehyde + Glycine Glycine + NAD + Tetrahydrofolic acid > Carbon dioxide + 5,10-Methylene-THF + NADH + Ammonium Adenosine triphosphate + Glycine + tRNA(Gly) + tRNA(Gly) <> Adenosine monophosphate + Glycyl-tRNA(Gly) + Pyrophosphate + Glycyl-tRNA(Gly) Water + L-Prolinylglycine > Glycine + L-Proline Water + Oxygen + Sarcosine > Formaldehyde + Glycine + Hydrogen peroxide L-Serine + Tetrahydrofolic acid <> Glycine + Water + 5,10-Methylene-THF Adenosine triphosphate + gamma-Glutamylcysteine + Glycine <> ADP + Glutathione + Hydrogen ion + Phosphate Acetyl-CoA + Glycine <> L-2-Amino-3-oxobutanoic acid + Coenzyme A Adenosine triphosphate + Glycine + 5-Phosphoribosylamine <> ADP + Glycineamideribotide + Hydrogen ion + Phosphate Adenosine triphosphate + gamma-Glutamylcysteine + Glycine <> ADP + Phosphate + Glutathione Glycine + Tetrahydrofolic acid + NAD <> 5,10-Methylene-THF + Ammonia + Carbon dioxide + NADH + Hydrogen ion Glycine + Lipoylprotein <> S-Aminomethyldihydrolipoylprotein + Carbon dioxide Adenosine triphosphate + Glycine + tRNA(Gly) <> Adenosine monophosphate + Pyrophosphate + Glycyl-tRNA(Gly) Adenosine triphosphate + 5-Phosphoribosylamine + Glycine <> ADP + Phosphate + Glycineamideribotide R-S-Cysteinylglycine + Water <> S-Substituted L-cysteine + Glycine L-Serine + 5,6,7,8-Tetrahydromethanopterin <> 5,10-Methylenetetrahydromethanopterin + Glycine + Water Glycine + Acetyl-CoA <> Hydrogen ion + L-2-Amino-3-oxobutanoic acid + Coenzyme A L-<i>threo</i>-3-phenylserine benzaldehyde + Glycine More...DL-allothreonine <> Acetaldehyde + Glycine 4-Hydroxy-L-threonine <> Glycolaldehyde + Glycine gly-met + Water > Glycine + L-Methionine ala-gly + Water > L-Alanine + Glycine gly-asn + Water > Glycine + L-Asparagine gly-gln + Water > Glycine + L-Glutamine glycyl-L-glutamate + Water > Glycine + L-Glutamate gly-asp + Water > Glycine + L-Aspartic acid glycylproline + Water > Glycine + L-Proline Glycine + H-protein-lipoyllysine > H-protein-S-aminomethyldihydrolipoyllysine + Carbon dioxide Adenosine triphosphate + gamma-Glutamylcysteine + Glycine > ADP + Inorganic phosphate + Glutathione Adenosine triphosphate + 5-Phosphoribosylamine + Glycine > ADP + Inorganic phosphate + 5'-Phospho-ribosylglycinamide Tetrahydrofolic acid + L-Serine + Tetrahydrofolic acid + L-Serine <> 5,10-Methylene-THF + Glycine + Water + 5,10-Methylene-THF L-Alanine + Glyoxylic acid + L-Alanine <> Glycine + Pyruvic acid Glycine + Adenosine triphosphate + Hydrogen ion + tRNA(gly) > Adenosine monophosphate + Pyrophosphate + Glycyl-tRNA(Gly) gamma-Glutamylcysteine + Glycine + Adenosine triphosphate > Hydrogen ion + Phosphate + Adenosine diphosphate + Glutathione + ADP 5-Phosphoribosylamine + Glycine + Adenosine triphosphate + 5-Phosphoribosylamine > N1-(5-phospho-β-D-ribosyl)glycinamide + Phosphate + Adenosine diphosphate + Hydrogen ion + ADP | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Pathways: |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Spectra | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Spectra: | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| References | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| References: |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Synthesis Reference: | Anslow, Winston K.; King, Harold. Synthesis of glycine. Journal of the Chemical Society (1929), 2163-6. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Material Safety Data Sheet (MSDS) | Download (PDF) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Links | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| External Links: |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
Enzymes
- General function:
- Involved in nucleotide binding
- Specific function:
- ATP + glycine + tRNA(Gly) = AMP + diphosphate + glycyl-tRNA(Gly)
- Gene Name:
- glyQ
- Locus Tag:
- PA0009
- Molecular weight:
- 36.1 kDa
Reactions
| ATP + glycine + tRNA(Gly) = AMP + diphosphate + glycyl-tRNA(Gly). |
- General function:
- Involved in arginine-tRNA ligase activity
- Specific function:
- ATP + glycine + tRNA(Gly) = AMP + diphosphate + glycyl-tRNA(Gly)
- Gene Name:
- glyS
- Locus Tag:
- PA0008
- Molecular weight:
- 74 kDa
Reactions
| ATP + glycine + tRNA(Gly) = AMP + diphosphate + glycyl-tRNA(Gly). |
- General function:
- Involved in ATP binding
- Specific function:
- ATP + gamma-L-glutamyl-L-cysteine + glycine = ADP + phosphate + glutathione
- Gene Name:
- gshB
- Locus Tag:
- PA0407
- Molecular weight:
- 35.7 kDa
Reactions
| ATP + gamma-L-glutamyl-L-cysteine + glycine = ADP + phosphate + glutathione. |
- General function:
- Involved in proteolysis
- Specific function:
- Aminopeptidase N is involved in the degradation of intracellular peptides generated by protein breakdown during normal growth as well as in response to nutrient starvation
- Gene Name:
- pepN
- Locus Tag:
- PA3083
- Molecular weight:
- 100 kDa
Reactions
| Release of an N-terminal amino acid, Xaa-|-Yaa- from a peptide, amide or arylamide. Xaa is preferably Ala, but may be most amino acids including Pro (slow action). When a terminal hydrophobic residue is followed by a prolyl residue, the two may be released as an intact Xaa-Pro dipeptide. |
- General function:
- Involved in catalytic activity
- Specific function:
- Interconversion of serine and glycine
- Gene Name:
- glyA
- Locus Tag:
- PA4602
- Molecular weight:
- 45.2 kDa
Reactions
| 5,10-methylenetetrahydrofolate + glycine + H(2)O = tetrahydrofolate + L-serine. |
- General function:
- Involved in ATP binding
- Specific function:
- ATP + 5-phospho-D-ribosylamine + glycine = ADP + phosphate + N(1)-(5-phospho-D-ribosyl)glycinamide
- Gene Name:
- purD
- Locus Tag:
- PA4855
- Molecular weight:
- 45.2 kDa
Reactions
| ATP + 5-phospho-D-ribosylamine + glycine = ADP + phosphate + N(1)-(5-phospho-D-ribosyl)glycinamide. |
- General function:
- Involved in aminomethyltransferase activity
- Specific function:
- The glycine cleavage system catalyzes the degradation of glycine
- Gene Name:
- gcvT
- Locus Tag:
- PA5215
- Molecular weight:
- 38.9 kDa
Reactions
| [Protein]-S(8)-aminomethyldihydrolipoyllysine + tetrahydrofolate = [protein]-dihydrolipoyllysine + 5,10-methylenetetrahydrofolate + NH(3). |
- General function:
- Involved in glycine dehydrogenase (decarboxylating) activity
- Specific function:
- The glycine cleavage system catalyzes the degradation of glycine. The P protein binds the alpha-amino group of glycine through its pyridoxal phosphate cofactor; CO(2) is released and the remaining methylamine moiety is then transferred to the lipoamide cofactor of the H protein
- Gene Name:
- gcvP
- Locus Tag:
- PA5213
- Molecular weight:
- 104.7 kDa
Reactions
| Glycine + H-protein-lipoyllysine = H-protein-S-aminomethyldihydrolipoyllysine + CO(2). |
- General function:
- Involved in aminopeptidase activity
- Specific function:
- Presumably involved in the processing and regular turnover of intracellular proteins. Catalyzes the removal of unsubstituted N-terminal amino acids from various peptides. Required for plasmid ColE1 site-specific recombination but not in its aminopeptidase activity. Could act as a structural component of the putative nucleoprotein complex in which the Xer recombination reaction takes place
- Gene Name:
- pepA
- Locus Tag:
- PA3831
- Molecular weight:
- 52.3 kDa
Reactions
| Release of an N-terminal amino acid, Xaa-|-Yaa-, in which Xaa is preferably Leu, but may be other amino acids including Pro although not Arg or Lys, and Yaa may be Pro. Amino acid amides and methyl esters are also readily hydrolyzed, but rates on arylamides are exceedingly low. |
| Release of an N-terminal amino acid, preferentially leucine, but not glutamic or aspartic acids. |
- General function:
- Involved in lyase activity
- Specific function:
- Catalyzes the cleavage of L-allo-threonine and L- threonine to glycine and acetaldehyde. L-threo-phenylserine and L- erythro-phenylserine are also good substrates
- Gene Name:
- ltaE
- Locus Tag:
- PA0902
- Molecular weight:
- 35.4 kDa
Reactions
| L-threonine = glycine + acetaldehyde. |
| L-allo-threonine = glycine + acetaldehyde. |
- General function:
- Amino acid transport and metabolism
- Specific function:
- The glycine cleavage system catalyzes the degradation of glycine. The H protein shuttles the methylamine group of glycine from the P protein to the T protein
- Gene Name:
- gcvH
- Locus Tag:
- PA5214
- Molecular weight:
- 13.6 kDa
Transporters
- General function:
- Involved in proteolysis
- Specific function:
- Aminopeptidase N is involved in the degradation of intracellular peptides generated by protein breakdown during normal growth as well as in response to nutrient starvation
- Gene Name:
- pepN
- Locus Tag:
- PA3083
- Molecular weight:
- 100 kDa
Reactions
| Release of an N-terminal amino acid, Xaa-|-Yaa- from a peptide, amide or arylamide. Xaa is preferably Ala, but may be most amino acids including Pro (slow action). When a terminal hydrophobic residue is followed by a prolyl residue, the two may be released as an intact Xaa-Pro dipeptide. |
- General function:
- Involved in sodium:amino acid symporter activity
- Specific function:
- Specific function unknown
- Gene Name:
- yaaJ
- Locus Tag:
- PA3641
- Molecular weight:
- 50.3 kDa
- General function:
- Involved in nucleotide binding
- Specific function:
- Probably part of a binding-protein-dependent transport system yecCS for an amino acid. Probably responsible for energy coupling to the transport system
- Gene Name:
- yecC
- Locus Tag:
- PA5152
- Molecular weight:
- 28.4 kDa