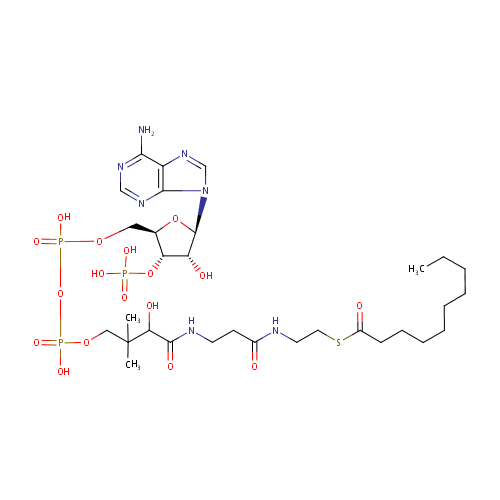
Decanoyl-CoA (N-C10:0CoA) (PAMDB000670)
| Record Information | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Version | 1.0 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Update Date | 1/22/2018 12:54:54 PM | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Metabolite ID | PAMDB000670 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Identification | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Name: | Decanoyl-CoA (N-C10:0CoA) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Description: | Decanoyl-CoA is an intermediate in fatty acid elongationand its formation is from acetyl-CoA. It is also involved in fatty acid metabolism. (KEGG) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Structure | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Synonyms: |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Chemical Formula: | C31H54N7O17P3S | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Average Molecular Weight: | 921.783 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Monoisotopic Molecular Weight: | 921.250973563 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| InChI Key: | CNKJPHSEFDPYDB-BOJFXZHGSA-N | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| InChI: | InChI=1S/C31H54N7O17P3S/c1-4-5-6-7-8-9-10-11-22(40)59-15-14-33-21(39)12-13-34-29(43)26(42)31(2,3)17-52-58(49,50)55-57(47,48)51-16-20-25(54-56(44,45)46)24(41)30(53-20)38-19-37-23-27(32)35-18-36-28(23)38/h18-20,24-26,30,41-42H,4-17H2,1-3H3,(H,33,39)(H,34,43)(H,47,48)(H,49,50)(H2,32,35,36)(H2,44,45,46)/t20-,24-,25-,26?,30-/m1/s1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| CAS number: | 1264-57-9 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| IUPAC Name: | {[(2R,3S,4R,5R)-5-(6-amino-9H-purin-9-yl)-2-[({[({3-[(2-{[2-(decanoylsulfanyl)ethyl]carbamoyl}ethyl)carbamoyl]-3-hydroxy-2,2-dimethylpropoxy}(hydroxy)phosphoryl)oxy](hydroxy)phosphoryl}oxy)methyl]-4-hydroxyoxolan-3-yl]oxy}phosphonic acid | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Traditional IUPAC Name: | [(2R,3S,4R,5R)-5-(6-aminopurin-9-yl)-2-{[({3-[(2-{[2-(decanoylsulfanyl)ethyl]carbamoyl}ethyl)carbamoyl]-3-hydroxy-2,2-dimethylpropoxy(hydroxy)phosphoryl}oxy(hydroxy)phosphoryl)oxy]methyl}-4-hydroxyoxolan-3-yl]oxyphosphonic acid | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| SMILES: | CCCCCCCCCC(=O)SCCNC(=O)CCNC(=O)C(O)C(C)(C)COP(O)(=O)OP(O)(=O)OC[C@H]1O[C@H]([C@H](O)[C@@H]1OP(O)(O)=O)N1C=NC2=C1N=CN=C2N | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Chemical Taxonomy | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Taxonomy Description | This compound belongs to the class of organic compounds known as 2,3,4-saturated fatty acyl coas. These are acyl-CoAs carrying a 2,3,4-saturated fatty acyl chain. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Kingdom | Organic compounds | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Super Class | Lipids and lipid-like molecules | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Class | Fatty Acyls | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Sub Class | Fatty acyl thioesters | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Direct Parent | 2,3,4-saturated fatty acyl CoAs | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Alternative Parents |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Substituents |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Molecular Framework | Aromatic heteropolycyclic compounds | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| External Descriptors |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Physical Properties | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| State: | Solid | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Charge: | -4 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Melting point: | Not Available | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Experimental Properties: |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Predicted Properties |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Biological Properties | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Cellular Locations: | Cytoplasm | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Reactions: | Adenosine triphosphate + Coenzyme A + Decanoate (N-C10:0) + Hydrogen ion > Adenosine monophosphate + Decanoyl-CoA (N-C10:0CoA) + Hydrogen ion + Pyrophosphate Acetyl-CoA + Decanoyl-CoA (N-C10:0CoA) <> 3-Oxododecanoyl-CoA + Coenzyme A Decanoyl-CoA (N-C10:0CoA) + FAD <> (2E)-Decenoyl-CoA + FADH2 Decanoyl-CoA (N-C10:0CoA) + Water > Coenzyme A + Decanoate (N-C10:0) + Hydrogen ion 3-Oxooctanoyl-CoA + Coenzyme A > Acetyl-CoA + Decanoyl-CoA (n-C10:0CoA) + Decanoyl-CoA (N-C10:0CoA) Decanoyl-CoA (n-C10:0CoA) + electron-transfer flavoprotein + Decanoyl-CoA (N-C10:0CoA) > (2E)-Decenoyl-CoA + Reduced electron-transfer flavoprotein Decanoyl-CoA (n-C10:0CoA) + Acetyl-CoA + Decanoyl-CoA (N-C10:0CoA) > 3-Oxododecanoyl-CoA + Coenzyme A Adenosine triphosphate + Coenzyme A + Capric acid > Adenosine monophosphate + Decanoyl-CoA (n-C10:0CoA) + Decanoyl-CoA (N-C10:0CoA) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Pathways: |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Spectra | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Spectra: |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| References | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| References: |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Synthesis Reference: | Not Available | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Material Safety Data Sheet (MSDS) | Download (PDF) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Links | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| External Links: |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
Enzymes
- General function:
- Involved in acetyl-CoA C-acyltransferase activity
- Specific function:
- Catalyzes the final step of fatty acid oxidation in which acetyl-CoA is released and the CoA ester of a fatty acid two carbons shorter is formed. Involved in the aerobic and anaerobic degradation of long-chain fatty acids
- Gene Name:
- fadA
- Locus Tag:
- PA3013
- Molecular weight:
- 41.6 kDa
Reactions
| Acyl-CoA + acetyl-CoA = CoA + 3-oxoacyl-CoA. |
- General function:
- Involved in catalytic activity
- Specific function:
- Catalyzes the esterification, concomitant with transport, of exogenous long-chain fatty acids into metabolically active CoA thioesters for subsequent degradation or incorporation into phospholipids
- Gene Name:
- fadD
- Locus Tag:
- PA3299
- Molecular weight:
- 61.7 kDa
Reactions
| ATP + a long-chain fatty acid + CoA = AMP + diphosphate + an acyl-CoA. |
- General function:
- Involved in acyl-CoA dehydrogenase activity
- Specific function:
- Catalyzes the dehydrogenation of acyl-CoA
- Gene Name:
- fadE
- Locus Tag:
- PA2815
- Molecular weight:
- 88.8 kDa
Reactions
| An acyl-CoA + FAD = a dehydrogenated acyl-CoA + FADH(2). |
- General function:
- Involved in acyl-CoA hydrolase activity
- Specific function:
- Can hydrolyze a broad range of acyl-CoA thioesters. Its physiological function is not known
- Gene Name:
- tesB
- Locus Tag:
- PA3942
- Molecular weight:
- 32.9 kDa