|
Record Information |
|---|
| Version |
1.0 |
|---|
| Update Date |
1/22/2018 12:54:54 PM |
|---|
|
Metabolite ID | PAMDB000516 |
|---|
|
Identification |
|---|
| Name: |
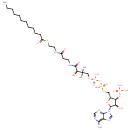
Lauroyl-CoA |
|---|
| Description: | Lauroyl-CoA is a substrate for Long chain fatty acyl-CoA synthetase. This enzyme is a member of the ligase family that activates the breakdown of complex fatty acids. Long chain fatty acyl-CoA synthetase plays a crucial role in intermediary metabolism by catalyzing the formation of fatty acyl-CoA by a two-step process proceeding through an adenylated intermediate. It is an enzyme present in all organisms from bacteria to humans. |
|---|
|
Structure |
|
|---|
| Synonyms: | - Dodecanoyl-CoA
- Dodecanoyl-CoA (N-C12:0CoA)
- Dodecanoyl-Coenzyme A
- Lauroyl coenzyme A
|
|---|
|
Chemical Formula: |
C33H58N7O17P3S |
|---|
| Average Molecular Weight: |
949.837 |
|---|
| Monoisotopic Molecular
Weight: |
949.282273691 |
|---|
| InChI Key: |
YMCXGHLSVALICC-GMHMEAMDSA-N |
|---|
| InChI: | InChI=1S/C33H58N7O17P3S/c1-4-5-6-7-8-9-10-11-12-13-24(42)61-17-16-35-23(41)14-15-36-31(45)28(44)33(2,3)19-54-60(51,52)57-59(49,50)53-18-22-27(56-58(46,47)48)26(43)32(55-22)40-21-39-25-29(34)37-20-38-30(25)40/h20-22,26-28,32,43-44H,4-19H2,1-3H3,(H,35,41)(H,36,45)(H,49,50)(H,51,52)(H2,34,37,38)(H2,46,47,48)/t22-,26-,27-,28+,32-/m1/s1 |
|---|
| CAS
number: |
6244-92-4 |
|---|
| IUPAC Name: | {[(2R,3S,4R,5R)-5-(6-amino-9H-purin-9-yl)-2-({[({[(3R)-3-[(2-{[2-(dodecanoylsulfanyl)ethyl]carbamoyl}ethyl)carbamoyl]-3-hydroxy-2,2-dimethylpropoxy](hydroxy)phosphoryl}oxy)(hydroxy)phosphoryl]oxy}methyl)-4-hydroxyoxolan-3-yl]oxy}phosphonic acid |
|---|
|
Traditional IUPAC Name: |
lauroyl-coa |
|---|
| SMILES: | CCCCCCCCCCCC(=O)SCCNC(=O)CCNC(=O)[C@H](O)C(C)(C)COP(O)(=O)OP(O)(=O)OC[C@H]1O[C@H]([C@H](O)[C@@H]1OP(O)(O)=O)N1C=NC2=C(N)N=CN=C12 |
|---|
|
Chemical Taxonomy |
|---|
|
Taxonomy Description | This compound belongs to the class of organic compounds known as 2,3,4-saturated fatty acyl coas. These are acyl-CoAs carrying a 2,3,4-saturated fatty acyl chain. |
|---|
|
Kingdom |
Organic compounds |
|---|
| Super Class | Lipids and lipid-like molecules |
|---|
|
Class |
Fatty Acyls |
|---|
| Sub Class | Fatty acyl thioesters |
|---|
|
Direct Parent |
2,3,4-saturated fatty acyl CoAs |
|---|
| Alternative Parents |
|
|---|
| Substituents |
- Coenzyme a or derivatives
- Purine ribonucleoside diphosphate
- Purine ribonucleoside 3',5'-bisphosphate
- N-glycosyl compound
- Glycosyl compound
- Beta amino acid or derivatives
- Organic pyrophosphate
- Monosaccharide phosphate
- 6-aminopurine
- Purine
- Imidazopyrimidine
- Monoalkyl phosphate
- Aminopyrimidine
- Imidolactam
- Alkyl phosphate
- Pyrimidine
- Primary aromatic amine
- Phosphoric acid ester
- Organic phosphoric acid derivative
- Organic phosphate
- N-substituted imidazole
- N-acyl-amine
- Monosaccharide
- Fatty amide
- Saccharide
- Heteroaromatic compound
- Oxolane
- Imidazole
- Azole
- Thiocarboxylic acid ester
- Secondary carboxylic acid amide
- Secondary alcohol
- Carboxamide group
- Oxacycle
- Azacycle
- Organoheterocyclic compound
- Sulfenyl compound
- Thioether
- Thiocarboxylic acid or derivatives
- Carboxylic acid derivative
- Carboxylic acid amide
- Hydrocarbon derivative
- Primary amine
- Organosulfur compound
- Organooxygen compound
- Organonitrogen compound
- Carbonyl group
- Amine
- Alcohol
- Aromatic heteropolycyclic compound
|
|---|
| Molecular Framework |
Aromatic heteropolycyclic compounds |
|---|
| External Descriptors |
|
|---|
|
Physical Properties |
|---|
| State: |
Solid |
|---|
| Charge: | -4 |
|---|
|
Melting point: |
Not Available |
|---|
| Experimental Properties: |
|
|---|
| Predicted Properties |
|
|---|
|
Biological Properties |
|---|
| Cellular Locations: |
Cytoplasm |
|---|
| Reactions: | |
|---|
|
Pathways: |
|
|---|
|
Spectra |
|---|
| Spectra: |
|
|---|
|
References |
|---|
| References: |
- Hunt MC, Ruiter J, Mooyer P, van Roermond CW, Ofman R, Ijlst L, Wanders RJ: Identification of fatty acid oxidation disorder patients with lowered acyl-CoA thioesterase activity in human skin fibroblasts. Eur J Clin Invest. 2005 Jan;35(1):38-46. Pubmed: 15638818
- Kanehisa, M., Goto, S., Sato, Y., Furumichi, M., Tanabe, M. (2012). "KEGG for integration and interpretation of large-scale molecular data sets." Nucleic Acids Res 40:D109-D114. Pubmed: 22080510
- Keseler, I. M., Collado-Vides, J., Santos-Zavaleta, A., Peralta-Gil, M., Gama-Castro, S., Muniz-Rascado, L., Bonavides-Martinez, C., Paley, S., Krummenacker, M., Altman, T., Kaipa, P., Spaulding, A., Pacheco, J., Latendresse, M., Fulcher, C., Sarker, M., Shearer, A. G., Mackie, A., Paulsen, I., Gunsalus, R. P., Karp, P. D. (2011). "EcoCyc: a comprehensive database of Escherichia coli biology." Nucleic Acids Res 39:D583-D590. Pubmed: 21097882
|
|---|
| Synthesis Reference: |
Eaton, Simon; Turnbull, Douglass M.; Bartlett, Kim. Redox control of b-oxidation in rat liver mitochondria. European Journal of Biochemistry (1994), 220(3), 671-81. |
|---|
| Material Safety Data Sheet (MSDS) |
Not Available |
|---|
|
Links |
|---|
| External Links: |
|
|---|