Phosphoadenosine phosphosulfate (PAMDB000269)
| Record Information | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Version | 1.0 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Update Date | 1/22/2018 12:54:54 PM | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Metabolite ID | PAMDB000269 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Identification | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
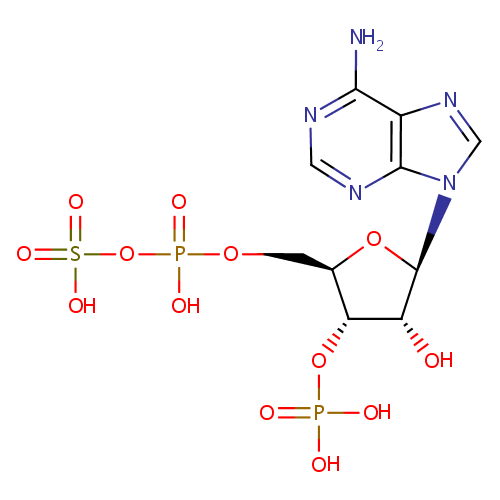
| Name: | Phosphoadenosine phosphosulfate | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Description: | 3'-Phosphoadenosine-5'-phosphosulfate is a key intermediate in the formation by living cells of sulfate esters of phenols, alcohols, sulfated polysaccharides, and simple esters, such as choline sulfate. It is formed from a sulfate ion and ATP in a two-step process. This compound also is an important intermediate in the process of sulfur fixation in plants and microorganisms. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Structure | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Synonyms: |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Chemical Formula: | C10H15N5O13P2S | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Average Molecular Weight: | 507.264 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Monoisotopic Molecular Weight: | 506.986229305 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| InChI Key: | GACDQMDRPRGCTN-KQYNXXCUSA-N | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| InChI: | InChI=1S/C10H15N5O13P2S/c11-8-5-9(13-2-12-8)15(3-14-5)10-6(16)7(27-29(17,18)19)4(26-10)1-25-30(20,21)28-31(22,23)24/h2-4,6-7,10,16H,1H2,(H,20,21)(H2,11,12,13)(H2,17,18,19)(H,22,23,24)/t4-,6-,7-,10-/m1/s1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| CAS number: | 482-67-7 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| IUPAC Name: | [({[(2R,3S,4R,5R)-5-(6-amino-9H-purin-9-yl)-4-hydroxy-3-(phosphonooxy)oxolan-2-yl]methoxy}(hydroxy)phosphoryl)oxy]sulfonic acid | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Traditional IUPAC Name: | 3'-phosphoadenylyl sulfate | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| SMILES: | NC1=NC=NC2=C1N=CN2[C@@H]1O[C@H](COP(O)(=O)OS(O)(=O)=O)[C@@H](OP(O)(O)=O)[C@H]1O | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Chemical Taxonomy | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Taxonomy Description | This compound belongs to the class of organic compounds known as purine ribonucleoside 3',5'-bisphosphates. These are purine ribobucleotides with one phosphate group attached to 3' and 5' hydroxyl groups of the ribose moiety. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Kingdom | Organic compounds | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Super Class | Nucleosides, nucleotides, and analogues | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Class | Purine nucleotides | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Sub Class | Purine ribonucleotides | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Direct Parent | Purine ribonucleoside 3',5'-bisphosphates | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Alternative Parents |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Substituents |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Molecular Framework | Aromatic heteropolycyclic compounds | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| External Descriptors |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Physical Properties | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| State: | Solid | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Charge: | -4 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Melting point: | Not Available | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Experimental Properties: |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Predicted Properties |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Biological Properties | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Cellular Locations: | Cytoplasm | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Reactions: | glutaredoxin + Phosphoadenosine phosphosulfate > glutaredoxin +2 Hydrogen ion + Adenosine 3',5'-diphosphate + Sulfite Phosphoadenosine phosphosulfate + Reduced Thioredoxin >2 Hydrogen ion + Adenosine 3',5'-diphosphate + Sulfite + Oxidized Thioredoxin Adenosine phosphosulfate + Adenosine triphosphate <> ADP + Hydrogen ion + Phosphoadenosine phosphosulfate Phosphoadenosine phosphosulfate + Water <> Adenosine phosphosulfate + Phosphate Adenosine triphosphate + Adenosine phosphosulfate <> ADP + Phosphoadenosine phosphosulfate Thioredoxin + Phosphoadenosine phosphosulfate + Thioredoxin disulfide <> Thioredoxin disulfide + Sulfite + Adenosine 3',5'-diphosphate + Thioredoxin Adenosine 3',5'-diphosphate + Sulfite + thioredoxin disulfide > Phosphoadenosine phosphosulfate + thioredoxin Phosphoadenosine phosphosulfate + reduced thioredoxin > Sulfite +2 Hydrogen ion + Adenosine 3',5'-diphosphate +2 oxidized thioredoxin + Sulfite + Adenosine 3',5'-diphosphate Phosphoadenosine phosphosulfate + reduced thioredoxin > Sulfite + oxidized thioredoxin + Hydrogen ion + Adenosine 3',5'-diphosphate + Sulfite + Adenosine 3',5'-diphosphate Adenosine phosphosulfate + Adenosine triphosphate > Phosphoadenosine phosphosulfate + Adenosine diphosphate + Hydrogen ion + ADP | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Pathways: | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Spectra | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Spectra: |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| References | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| References: |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Synthesis Reference: | Lin, Chun-Hung; Shen, Gwo-Jenn; Garcia-Junceda, Eduardo; Wong, Chi-Huey. Enzymic Synthesis and Regeneration of 3'-Phosphoadenosine 5'-Phosphosulfate (PAPS) for Regioselective Sulfation of Oligosaccharides. Journal of the American Chemical Society (1995), | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Material Safety Data Sheet (MSDS) | Not Available | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Links | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| External Links: |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
Enzymes
- General function:
- Involved in adenylylsulfate kinase activity
- Specific function:
- Catalyzes the synthesis of activated sulfate
- Gene Name:
- cysC
- Locus Tag:
- PA1393
- Molecular weight:
- 22.1 kDa
Reactions
| ATP + adenylyl sulfate = ADP + 3'-phosphoadenylyl sulfate. |
- General function:
- Involved in phosphoadenylyl-sulfate reductase (thioredoxin) activity
- Specific function:
- Reduction of activated sulfate into sulfite
- Gene Name:
- cysH
- Locus Tag:
- PA1756
- Molecular weight:
- 30.2 kDa
Reactions
| Adenosine 3',5'-bisphosphate + sulfite + thioredoxin disulfide = 3'-phosphoadenylyl sulfate + thioredoxin. |
- General function:
- Involved in magnesium ion binding
- Specific function:
- Converts 3'(2')-phosphoadenosine 5'-phosphate (PAP) to AMP. May also convert adenosine 3'-phosphate 5'-phosphosulfate (PAPS) to adenosine 5'-phosphosulfate (APS). Has 10000-fold lower activity towards inositol 1,4-bisphosphate (Ins(1,4)P2)
- Gene Name:
- cysQ
- Locus Tag:
- PA5175
- Molecular weight:
- 29.8 kDa
Reactions
| Adenosine 3',5'-bisphosphate + H(2)O = adenosine 5'-phosphate + phosphate. |
- General function:
- Involved in transporter activity
- Specific function:
- Part of a binding-protein-dependent transport system for aliphatic sulfonates. Probably responsible for the translocation of the substrate across the membrane
- Gene Name:
- ssuC
- Locus Tag:
- PA3443
- Molecular weight:
- 28.5 kDa
- General function:
- Involved in electron carrier activity
- Specific function:
- Monothiol glutaredoxin involved in the biogenesis of iron-sulfur clusters (Probable)
- Gene Name:
- grxD
- Locus Tag:
- PA3533
- Molecular weight:
- 11.8 kDa
- General function:
- Involved in electron carrier activity
- Specific function:
- The disulfide bond functions as an electron carrier in the glutathione-dependent synthesis of deoxyribonucleotides by the enzyme ribonucleotide reductase. In addition, it is also involved in reducing some disulfides in a coupled system with glutathione reductase
- Gene Name:
- grxC
- Locus Tag:
- PA5129
- Molecular weight:
- 9.2 kDa
- General function:
- Involved in electron carrier activity
- Specific function:
- Participates in various redox reactions through the reversible oxidation of its active center dithiol to a disulfide and catalyzes dithiol-disulfide exchange reactions
- Gene Name:
- trxA
- Locus Tag:
- PA5240
- Molecular weight:
- 11.9 kDa
Transporters
- General function:
- Involved in transporter activity
- Specific function:
- Part of a binding-protein-dependent transport system for aliphatic sulfonates. Probably responsible for the translocation of the substrate across the membrane
- Gene Name:
- ssuC
- Locus Tag:
- PA3443
- Molecular weight:
- 28.5 kDa