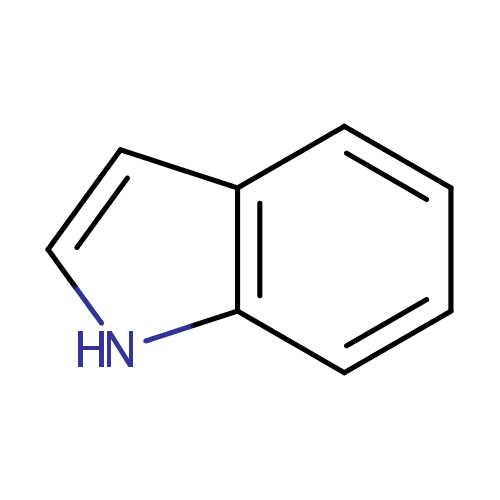
Indole (PAMDB000181)
| Record Information | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Version | 1.0 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Update Date | 1/22/2018 12:54:54 PM | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Metabolite ID | PAMDB000181 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Identification | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Name: | Indole | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Description: | Indole is an aromatic heterocyclic organic compound. It has a bicyclic structure, consisting of a six-membered benzene ring fused to a five-membered nitrogen-containing pyrrole ring. It can be produced by bacteria as a degradation product of the amino acid tryptophan. It occurs naturally in feces and has an intense fecal smell. At very low concentrations, however, it has a flowery smell, and is a constituent of many flower scents (such as orange blossoms) and perfumes. Natural jasmine oil, used in the perfume industry, contains around 2.5% of indole. Indole also occurs in CoAl tar. The participation of the nitrogen lone electron pair in the aromatic ring means that indole is not a base, and it does not behave like a simple amine. | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Structure | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Synonyms: |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Chemical Formula: | C8H7N | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Average Molecular Weight: | 117.1479 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Monoisotopic Molecular Weight: | 117.057849229 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| InChI Key: | SIKJAQJRHWYJAI-UHFFFAOYSA-N | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| InChI: | InChI=1S/C8H7N/c1-2-4-8-7(3-1)5-6-9-8/h1-6,9H | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| CAS number: | 120-72-9 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| IUPAC Name: | 1H-indole | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Traditional IUPAC Name: | indole | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| SMILES: | N1C=CC2=C1C=CC=C2 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Chemical Taxonomy | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Taxonomy Description | This compound belongs to the class of organic compounds known as indoles. These are compounds containing an indole moiety, which consists of pyrrole ring fused to benzene to form 2,3-benzopyrrole. | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Kingdom | Organic compounds | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Super Class | Organoheterocyclic compounds | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Class | Indoles and derivatives | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Sub Class | Indoles | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Direct Parent | Indoles | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Alternative Parents | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Substituents |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Molecular Framework | Aromatic heteropolycyclic compounds | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| External Descriptors |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Physical Properties | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| State: | Solid | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Charge: | 0 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Melting point: | 52.5 °C | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Experimental Properties: |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Predicted Properties |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Biological Properties | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Cellular Locations: | Cytoplasm | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Reactions: | Indole + L-Serine > Water + L-Tryptophan Indoleglycerol phosphate > D-Glyceraldehyde 3-phosphate + Indole Water + L-Tryptophan <> Indole + Ammonium + Pyruvic acid L-Tryptophan + Water <> Indole + Pyruvic acid + Ammonia L-Tryptophan + Water <> Hydrogen ion + Indole + Pyruvic acid + Ammonia L-Tryptophan + Water + 2-Aminoacrylic acid + 2-Iminopropanoate <> Indole + Pyruvic acid + Ammonia L-Serine + Indoleglycerol phosphate + Indole <> L-Tryptophan + D-Glyceraldehyde 3-phosphate + Water (1S,2R)-1-C-(indol-3-yl)glycerol 3-phosphate + (1S,2R)-1-C-(indol-3-yl)glycerol 3-phosphate > D-Glyceraldehyde 3-phosphate + Indole + D-Glyceraldehyde 3-phosphate Indole + L-Serine + L-Serine > Water + L-Tryptophan L-Tryptophan > Hydrogen ion + Indole + 2-Aminoacrylic acid | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Pathways: | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Spectra | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Spectra: | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| References | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| References: |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Synthesis Reference: | Grigoleit, Georg; Oberkobusch, Rudolf; Collin, Gerd. Indole from 2-ethylaniline. Ger. Offen. (1973), 6 pp. | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Material Safety Data Sheet (MSDS) | Download (PDF) | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Links | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| External Links: |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||
Enzymes
- General function:
- Involved in catalytic activity
- Specific function:
- The alpha subunit is responsible for the aldol cleavage of indoleglycerol phosphate to indole and glyceraldehyde 3- phosphate
- Gene Name:
- trpA
- Locus Tag:
- PA0035
- Molecular weight:
- 28.5 kDa
Reactions
| L-serine + 1-C-(indol-3-yl)glycerol 3-phosphate = L-tryptophan + glyceraldehyde 3-phosphate + H(2)O. |
- General function:
- Involved in catalytic activity
- Specific function:
- The beta subunit is responsible for the synthesis of L- tryptophan from indole and L-serine
- Gene Name:
- trpB
- Locus Tag:
- PA0036
- Molecular weight:
- 43.7 kDa
Reactions
| L-serine + 1-C-(indol-3-yl)glycerol 3-phosphate = L-tryptophan + glyceraldehyde 3-phosphate + H(2)O. |
Transporters
- General function:
- Involved in amino acid transmembrane transporter activity
- Specific function:
- Involved in transporting tryptophan across the cytoplasmic membrane
- Gene Name:
- mtr
- Locus Tag:
- PA5434
- Molecular weight:
- 44.7 kDa