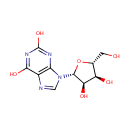
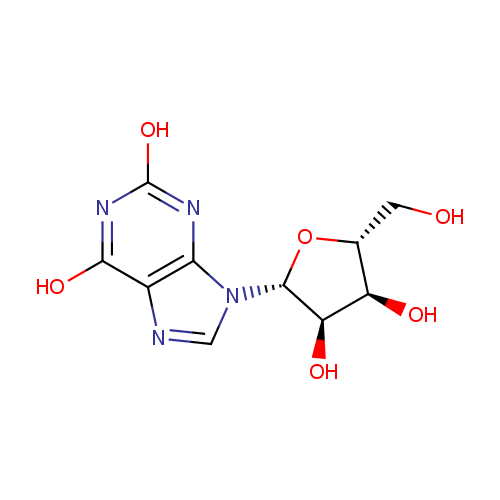
| Synonyms: | - 3,9-Dihydro-9-b-D-ribofuranosyl-1H-purine-2,6-dione
- 3,9-dihydro-9-b-delta-Ribofuranosyl-1H-purine-2,6-dione
- 3,9-dihydro-9-b-δ-Ribofuranosyl-1H-purine-2,6-dione
- 3,9-Dihydro-9-beta-delta-ribofuranosyl-1H-purine-2,6-dione
- 3,9-Dihydro-9-D-ribofuranosyl-1H-purine-2,6-dione
- 3,9-Dihydro-9-delta-ribofuranosyl-1H-purine-2,6-dione
- 3,9-dihydro-9-β-δ-Ribofuranosyl-1H-purine-2,6-dione
- 3,9-dihydro-9-δ-Ribofuranosyl-1H-purine-2,6-dione
- 9 β-D-ribofuranosylxanthine
- 9 b-D-Ribofuranosylxanthine
- 9 beta-D-Ribofuranosylxanthine
- 9 β-D-Ribofuranosylxanthine
- 9-b-D-Ribofuranosylxanthine
- 9-b-delta-Ribofuranosylxanthine
- 9-b-δ-Ribofuranosylxanthine
- 9-beta-delta-Ribofuranosylxanthine
- 9-D-Ribofuranosylxanthine
- 9-delta-Ribofuranosylxanthine
- 9-β-δ-Ribofuranosylxanthine
- 9-δ-Ribofuranosylxanthine
- Xanthosine
|
|---|
| References: |
- Dudley E, Lemiere F, Van Dongen W, Tuytten R, El-Sharkawi S, Brenton AG, Esmans EL, Newton RP: Analysis of urinary nucleosides. IV. Identification of urinary purine nucleosides by liquid chromatography/electrospray mass spectrometry. Rapid Commun Mass Spectrom. 2004;18(22):2730-8. Pubmed: 15499664
- Kanehisa, M., Goto, S., Sato, Y., Furumichi, M., Tanabe, M. (2012). "KEGG for integration and interpretation of large-scale molecular data sets." Nucleic Acids Res 40:D109-D114. Pubmed: 22080510
- Keseler, I. M., Collado-Vides, J., Santos-Zavaleta, A., Peralta-Gil, M., Gama-Castro, S., Muniz-Rascado, L., Bonavides-Martinez, C., Paley, S., Krummenacker, M., Altman, T., Kaipa, P., Spaulding, A., Pacheco, J., Latendresse, M., Fulcher, C., Sarker, M., Shearer, A. G., Mackie, A., Paulsen, I., Gunsalus, R. P., Karp, P. D. (2011). "EcoCyc: a comprehensive database of Escherichia coli biology." Nucleic Acids Res 39:D583-D590. Pubmed: 21097882
- Khalil PN, Erb N, Khalil MN, Escherich G, Janka-Schaub GE: Validation and application of a high-performance liquid chromatographic-based assay for determination of the inosine 5'-monophosphate dehydrogenase activity in erythrocytes. J Chromatogr B Analyt Technol Biomed Life Sci. 2006 Sep 14;842(1):1-7. Epub 2006 May 24. Pubmed: 16725387
- Liebich HM, Di Stefano C, Wixforth A, Schmid HR: Quantitation of urinary nucleosides by high-performance liquid chromatography. J Chromatogr A. 1997 Feb 28;763(1-2):193-7. Pubmed: 9129323
- Marti R, Nishigaki Y, Hirano M: Elevated plasma deoxyuridine in patients with thymidine phosphorylase deficiency. Biochem Biophys Res Commun. 2003 Mar 28;303(1):14-8. Pubmed: 12646159
- van der Werf, M. J., Overkamp, K. M., Muilwijk, B., Coulier, L., Hankemeier, T. (2007). "Microbial metabolomics: toward a platform with full metabolome coverage." Anal Biochem 370:17-25. Pubmed: 17765195
- Winder, C. L., Dunn, W. B., Schuler, S., Broadhurst, D., Jarvis, R., Stephens, G. M., Goodacre, R. (2008). "Global metabolic profiling of Escherichia coli cultures: an evaluation of methods for quenching and extraction of intracellular metabolites." Anal Chem 80:2939-2948. Pubmed: 18331064
|
|---|