Adenosine 3',5'-diphosphate (PAMDB000022)
| Record Information | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Version | 1.0 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Update Date | 1/22/2018 12:54:54 PM | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Metabolite ID | PAMDB000022 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Identification | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
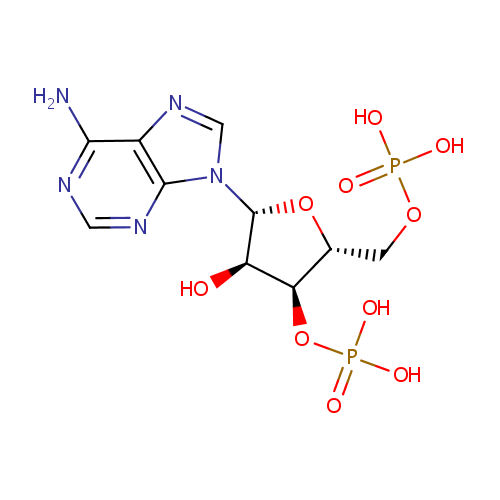
| Name: | Adenosine 3',5'-diphosphate | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Description: | Adenosine 3', 5'-diphosphate or PAP is a nucleotide that is closely related to ADP. It has two phosphate groups attached to the 5' and 3' positions of the pentose sugar ribose (instead of pyrophosphoric acid at the 5' position, as found in ADP), and the nucleobase adenine. PAP is converted to 3'-phosphoadenosine 5'-phosphosulfate (PAPS) by Sulfotransferase and then back to PAP after the sulfotransferase reaction. Sulfotransferase (STs) catalyze the transfer reaction of the sulfate group from the ubiquitous donor 3'-phosphoadenosine 5'-phosphosulfate (PAPS) to an acceptor group of numerous substrates. This reaction, often referred to as sulfuryl transfer, sulfation, or sulfonation plays a key role in various biological processes such as cell communication, growth and development, and defense. (HMDB) PAP is involved in sulfur metabolism in Pseudomonas aeruginosa, and can be converted into AMP and subsequently ADP by enzymes. (PMID 10939241) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Structure | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Synonyms: |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Chemical Formula: | C10H15N5O10P2 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Average Molecular Weight: | 427.2011 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Monoisotopic Molecular Weight: | 427.029414749 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| InChI Key: | WHTCPDAXWFLDIH-KQYNXXCUSA-N | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| InChI: | InChI=1S/C10H15N5O10P2/c11-8-5-9(13-2-12-8)15(3-14-5)10-6(16)7(25-27(20,21)22)4(24-10)1-23-26(17,18)19/h2-4,6-7,10,16H,1H2,(H2,11,12,13)(H2,17,18,19)(H2,20,21,22)/t4-,6-,7-,10-/m1/s1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| CAS number: | 1053-73-2 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| IUPAC Name: | {[(2R,3S,4R,5R)-5-(6-amino-9H-purin-9-yl)-4-hydroxy-3-(phosphonooxy)oxolan-2-yl]methoxy}phosphonic acid | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Traditional IUPAC Name: | adenosine 3',5'-bisphosphate | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| SMILES: | NC1=NC=NC2=C1N=CN2[C@@H]1O[C@H](COP(O)(O)=O)[C@@H](OP(O)(O)=O)[C@H]1O | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Chemical Taxonomy | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Taxonomy Description | This compound belongs to the class of organic compounds known as purine ribonucleoside 3',5'-bisphosphates. These are purine ribobucleotides with one phosphate group attached to 3' and 5' hydroxyl groups of the ribose moiety. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Kingdom | Organic compounds | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Super Class | Nucleosides, nucleotides, and analogues | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Class | Purine nucleotides | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Sub Class | Purine ribonucleotides | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Direct Parent | Purine ribonucleoside 3',5'-bisphosphates | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Alternative Parents |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Substituents |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Molecular Framework | Aromatic heteropolycyclic compounds | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| External Descriptors |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Physical Properties | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| State: | Solid | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Charge: | -4 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Melting point: | Not Available | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Experimental Properties: |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Predicted Properties |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Biological Properties | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Cellular Locations: | Cytoplasm | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Reactions: | glutaredoxin + Phosphoadenosine phosphosulfate > glutaredoxin +2 Hydrogen ion + Adenosine 3',5'-diphosphate + Sulfite Phosphoadenosine phosphosulfate + Reduced Thioredoxin >2 Hydrogen ion + Adenosine 3',5'-diphosphate + Sulfite + Oxidized Thioredoxin apoprotein [acyl carrier protein] + Coenzyme A > acyl carrier protein + Hydrogen ion + Adenosine 3',5'-diphosphate Water + Adenosine 3',5'-diphosphate > Adenosine monophosphate + Phosphate Coenzyme A + Apo-[acyl-carrier-protein] <> Adenosine 3',5'-diphosphate + Acyl-carrier protein + Acyl-carrier protein Thioredoxin + Phosphoadenosine phosphosulfate + Thioredoxin disulfide <> Thioredoxin disulfide + Sulfite + Adenosine 3',5'-diphosphate + Thioredoxin CoA-(4'-phosphopantetheine) + apo-[acyl-carrier-protein] > Adenosine 3',5'-diphosphate + holo-[acyl-carrier-protein] Adenosine 3',5'-diphosphate + Sulfite + thioredoxin disulfide > Phosphoadenosine phosphosulfate + thioredoxin Adenosine 3',5'-diphosphate + Water > Adenosine monophosphate + Inorganic phosphate Phosphoadenosine phosphosulfate + reduced thioredoxin > Sulfite +2 Hydrogen ion + Adenosine 3',5'-diphosphate +2 oxidized thioredoxin + Sulfite + Adenosine 3',5'-diphosphate Phosphoadenosine phosphosulfate + reduced thioredoxin > Sulfite + oxidized thioredoxin + Hydrogen ion + Adenosine 3',5'-diphosphate + Sulfite + Adenosine 3',5'-diphosphate | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Pathways: | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Spectra | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Spectra: |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| References | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| References: |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Synthesis Reference: | Tsunako, Mitsutomo; Kotone, Akira. Preparation of nucleoside-2',5'- and 3',5'-diphosphoric acids. Jpn. Kokai Tokkyo Koho (1991), 13 pp. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Material Safety Data Sheet (MSDS) | Download (PDF) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Links | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| External Links: |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
Enzymes
- General function:
- Involved in phosphoadenylyl-sulfate reductase (thioredoxin) activity
- Specific function:
- Reduction of activated sulfate into sulfite
- Gene Name:
- cysH
- Locus Tag:
- PA1756
- Molecular weight:
- 30.2 kDa
Reactions
| Adenosine 3',5'-bisphosphate + sulfite + thioredoxin disulfide = 3'-phosphoadenylyl sulfate + thioredoxin. |
- General function:
- Involved in magnesium ion binding
- Specific function:
- Converts 3'(2')-phosphoadenosine 5'-phosphate (PAP) to AMP. May also convert adenosine 3'-phosphate 5'-phosphosulfate (PAPS) to adenosine 5'-phosphosulfate (APS). Has 10000-fold lower activity towards inositol 1,4-bisphosphate (Ins(1,4)P2)
- Gene Name:
- cysQ
- Locus Tag:
- PA5175
- Molecular weight:
- 29.8 kDa
Reactions
| Adenosine 3',5'-bisphosphate + H(2)O = adenosine 5'-phosphate + phosphate. |
- General function:
- Involved in electron carrier activity
- Specific function:
- Monothiol glutaredoxin involved in the biogenesis of iron-sulfur clusters (Probable)
- Gene Name:
- grxD
- Locus Tag:
- PA3533
- Molecular weight:
- 11.8 kDa
- General function:
- Involved in electron carrier activity
- Specific function:
- The disulfide bond functions as an electron carrier in the glutathione-dependent synthesis of deoxyribonucleotides by the enzyme ribonucleotide reductase. In addition, it is also involved in reducing some disulfides in a coupled system with glutathione reductase
- Gene Name:
- grxC
- Locus Tag:
- PA5129
- Molecular weight:
- 9.2 kDa
- General function:
- Involved in electron carrier activity
- Specific function:
- Participates in various redox reactions through the reversible oxidation of its active center dithiol to a disulfide and catalyzes dithiol-disulfide exchange reactions
- Gene Name:
- trxA
- Locus Tag:
- PA5240
- Molecular weight:
- 11.9 kDa