Propionyl-CoA (PAMDB000321)
| Record Information | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Version | 1.0 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Update Date | 1/22/2018 12:54:54 PM | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Metabolite ID | PAMDB000321 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Identification | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
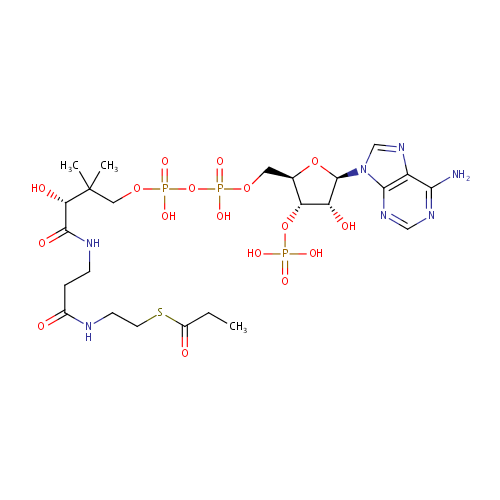
| Name: | Propionyl-CoA | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Description: | Propionyl-CoA is an intermediate in the metabolism of propanoate. Propionyl-CoA is a substrate for acetyl-CoA synthetase, propionyl-CoA synthetase and 2-methylcitrate synthase. It is also involved in beta-alanine metabolism, Valine, leucine abd isoleucine degradation and C5-branched dibasic acid metabolism pathways. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Structure | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Synonyms: |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Chemical Formula: | C24H40N7O17P3S | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Average Molecular Weight: | 823.597 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Monoisotopic Molecular Weight: | 823.141423115 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| InChI Key: | QAQREVBBADEHPA-UXYNFSPESA-N | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| InChI: | InChI=1S/C24H40N7O17P3S/c1-4-15(33)52-8-7-26-14(32)5-6-27-22(36)19(35)24(2,3)10-45-51(42,43)48-50(40,41)44-9-13-18(47-49(37,38)39)17(34)23(46-13)31-12-30-16-20(25)28-11-29-21(16)31/h11-13,17-19,23,34-35H,4-10H2,1-3H3,(H,26,32)(H,27,36)(H,40,41)(H,42,43)(H2,25,28,29)(H2,37,38,39)/t13-,17-,18-,19?,23-/m1/s1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| CAS number: | 317-66-8 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| IUPAC Name: | {[(2R,3S,4R,5R)-5-(6-amino-9H-purin-9-yl)-4-hydroxy-2-({[hydroxy({hydroxy[(3R)-3-hydroxy-2,2-dimethyl-3-[(2-{[2-(propanoylsulfanyl)ethyl]carbamoyl}ethyl)carbamoyl]propoxy]phosphoryl}oxy)phosphoryl]oxy}methyl)oxolan-3-yl]oxy}phosphonic acid | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Traditional IUPAC Name: | propionyl-coa | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| SMILES: | CCC(=O)SCCNC(=O)CCNC(=O)C(O)C(C)(C)COP(O)(=O)OP(O)(=O)OC[C@H]1O[C@H]([C@H](O)[C@@H]1OP(O)(O)=O)N1C=NC2=C1N=CN=C2N | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Chemical Taxonomy | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Taxonomy Description | This compound belongs to the class of organic compounds known as acyl coas. These are organic compounds containing a coenzyme A substructure linked to an acyl chain. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Kingdom | Organic compounds | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Super Class | Lipids and lipid-like molecules | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Class | Fatty Acyls | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Sub Class | Fatty acyl thioesters | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Direct Parent | Acyl CoAs | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Alternative Parents |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Substituents |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Molecular Framework | Aromatic heteropolycyclic compounds | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| External Descriptors |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Physical Properties | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| State: | Solid | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Charge: | -4 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Melting point: | Not Available | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Experimental Properties: |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Predicted Properties |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Biological Properties | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Cellular Locations: | Cytoplasm | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Reactions: | 2-Ketobutyric acid + Coenzyme A > Formic acid + Propionyl-CoA Water + Oxalacetic acid + Propionyl-CoA <> Methylcitric acid + Coenzyme A + Hydrogen ion + (2S,3S)-2-hydroxybutane-1,2,3-tricarboxylate Hydrogen ion + (S)-Methylmalonyl-CoA <> Carbon dioxide + Propionyl-CoA Propionyl-CoA + Succinic acid > Propionic acid + Succinyl-CoA Adenosine triphosphate + Coenzyme A + Propionic acid > ADP + Phosphate + Propionyl-CoA (S)-Methylmalonyl-CoA <> Propionyl-CoA + Carbon dioxide Propionyl-CoA + Acetyl-CoA <> Coenzyme A + 2-Methylacetoacetyl-CoA Methylcitric acid + Coenzyme A <> Propionyl-CoA + Oxalacetic acid + Water Propionyl-CoA + Chenodeoxycholoyl-CoA <> Coenzyme A + 3alpha,7alpha-Dihydroxy-5beta-cholestanoyl-CoA Oxalacetic acid + Water + Propionyl-CoA <> Hydrogen ion + Methylcitric acid + Coenzyme A Propionyl-CoA + Water + Glyoxylic acid <> 2-hydroxyglutarate + Hydrogen ion + Coenzyme A Coenzyme A + Propionic acid + Adenosine triphosphate > Propionyl-CoA + Pyrophosphate + Adenosine monophosphate Adenosine triphosphate + Hydrogen carbonate + Propionyl-CoA > Hydrogen ion + ADP + Phosphate + (S)-Methylmalonyl-CoA Propionyl-CoA + Phosphate > Propanoyl phosphate + Coenzyme A Hydrogen ion + R-Methylmalonyl-CoA > Propionyl-CoA + Carbon dioxide Propionyl-CoA + NADP <> Acrylyl-CoA + NADPH + Hydrogen ion More...2-Ketobutyric acid + Coenzyme A > Formic acid + Propionyl-CoA + Propionyl-CoA Propionyl-CoA + Phosphate + Propionyl-CoA > Coenzyme A + propanoyl phosphate + Propanoyl phosphate Propionic acid + Adenosine triphosphate + Coenzyme A > Propionyl-CoA + Adenosine monophosphate + Pyrophosphate + Propionyl-CoA Propionyl-CoA + Water + Oxalacetic acid + Propionyl-CoA > Coenzyme A + Hydrogen ion + 2-Methylcitric acid + Methylcitric acid Succinic acid + Propionyl-CoA + Propionyl-CoA > Propionic acid + Succinyl-CoA + Succinyl-CoA Methylmalonyl-CoA + Hydrogen ion > Carbon dioxide + Propionyl-CoA + Propionyl-CoA | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Pathways: |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Spectra | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Spectra: |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| References | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| References: |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Synthesis Reference: | Sokatch, John R.; Sanders, Lois E.; Marshall, Vincent P. Oxidation of methylmalonate semialdehyde to propionyl coenzyme A in Pseudomonas aeruginosa grown on valine. Journal of Biological Chemistry (1968), 243(10), 2500-6. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Material Safety Data Sheet (MSDS) | Not Available | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Links | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| External Links: |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
Enzymes
- General function:
- Involved in phosphate acetyltransferase activity
- Specific function:
- Acetyl-CoA + phosphate = CoA + acetyl phosphate
- Gene Name:
- pta
- Locus Tag:
- PA0835
- Molecular weight:
- 75.7 kDa
Reactions
| Acetyl-CoA + phosphate = CoA + acetyl phosphate. |
- General function:
- Involved in acetyl-CoA C-acyltransferase activity
- Specific function:
- Catalyzes the final step of fatty acid oxidation in which acetyl-CoA is released and the CoA ester of a fatty acid two carbons shorter is formed. Involved in the aerobic and anaerobic degradation of long-chain fatty acids
- Gene Name:
- fadA
- Locus Tag:
- PA3013
- Molecular weight:
- 41.6 kDa
Reactions
| Acyl-CoA + acetyl-CoA = CoA + 3-oxoacyl-CoA. |
- General function:
- Involved in acetate-CoA ligase activity
- Specific function:
- Enables the cell to use acetate during aerobic growth to generate energy via the TCA cycle, and biosynthetic compounds via the glyoxylate shunt. Acetylates CheY, the response regulator involved in flagellar movement and chemotaxis
- Gene Name:
- acs
- Locus Tag:
- PA0887
- Molecular weight:
- 71.8 kDa
Reactions
| ATP + acetate + CoA = AMP + diphosphate + acetyl-CoA. |
- General function:
- Involved in transferase activity, transferring acyl groups, acyl groups converted into alkyl on transfer
- Specific function:
- Catalyzes the synthesis of 2-methylcitrate from propionyl-CoA and oxaloacetate. Also catalyzes the condensation of oxaloacetate with acetyl-CoA but with a lower specificity
- Gene Name:
- prpC
- Locus Tag:
- PA0795
- Molecular weight:
- 41.7 kDa
Reactions
| Propanoyl-CoA + H(2)O + oxaloacetate = (2R,3S)-2-hydroxybutane-1,2,3-tricarboxylate + CoA. |