Ornithine (PAMDB000087)
| Record Information | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Version | 1.0 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Update Date | 1/22/2018 12:54:54 PM | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Metabolite ID | PAMDB000087 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Identification | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
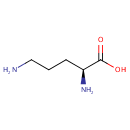
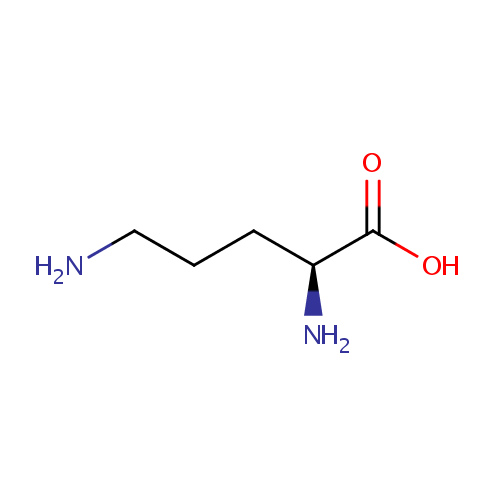
| Name: | Ornithine | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Description: | Ornithine is an amino acid produced in the urea cycle by the splitting off of urea from arginine. It is a central part of the urea cycle, which allows for the disposal of excess nitrogen. L-Ornithine is also a precursor of citrulline and arginine. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Structure | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Synonyms: |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Chemical Formula: | C5H12N2O2 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Average Molecular Weight: | 132.161 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Monoisotopic Molecular Weight: | 132.089877638 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| InChI Key: | AHLPHDHHMVZTML-BYPYZUCNSA-N | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| InChI: | InChI=1S/C5H12N2O2/c6-3-1-2-4(7)5(8)9/h4H,1-3,6-7H2,(H,8,9)/t4-/m0/s1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| CAS number: | 70-26-8 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| IUPAC Name: | (2S)-2,5-diaminopentanoic acid | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Traditional IUPAC Name: | ornithine | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| SMILES: | NCCC[C@H](N)C(O)=O | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Chemical Taxonomy | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Taxonomy Description | This compound belongs to the class of organic compounds known as d-alpha-amino acids. These are alpha amino acids which have the D-configuration of the alpha-carbon atom. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Kingdom | Organic compounds | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Super Class | Organic acids and derivatives | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Class | Carboxylic acids and derivatives | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Sub Class | Amino acids, peptides, and analogues | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Direct Parent | D-alpha-amino acids | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Alternative Parents | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Substituents |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Molecular Framework | Aliphatic acyclic compounds | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| External Descriptors |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Physical Properties | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| State: | Solid | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Charge: | 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Melting point: | 140 °C | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Experimental Properties: |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Predicted Properties |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Biological Properties | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Cellular Locations: | Cytoplasm | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Reactions: | Adenosine triphosphate + Water + Ornithine > ADP + Hydrogen ion + Ornithine + Phosphate Adenosine triphosphate + Water + Ornithine > ADP + Hydrogen ion + Ornithine + Phosphate Hydrogen ion + Ornithine + L-Ornithine <> Carbon dioxide + Putrescine + Ethylenediamine Carbamoylphosphate + Ornithine + L-Ornithine <> Citrulline + Hydrogen ion + Phosphate N-Acetylornithine + Water <> Acetic acid + Ornithine + L-Ornithine N-Acetylornithine + Water <> Acetic acid + Ornithine Ornithine <> Putrescine + Carbon dioxide Carbamoylphosphate + Ornithine <> Phosphate + Citrulline -->-->Ornithine + Carbamoylphosphate <> Hydrogen ion + Citrulline + Phosphate Hydrogen ion + Ornithine > Carbon dioxide + Putrescine Carbamoylphosphate + Ornithine > Inorganic phosphate + Citrulline N-Acetylornithine + Water > Ornithine + Acetic acid + Ornithine Ornithine + Carbamoylphosphate + Ornithine > Phosphate + Hydrogen ion + Citrulline Ornithine + Hydrogen ion + Ornithine > Putrescine + Carbon dioxide | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Pathways: | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Spectra | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Spectra: | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| References | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| References: |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Synthesis Reference: | Zhang, Peng; Zhang, Shurong; Liu, Chunqiao; Yang, Yuhong. Method for preparing L-ornithine by enzymatic conversion. Faming Zhuanli Shenqing Gongkai Shuomingshu (2007), 8pp. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Material Safety Data Sheet (MSDS) | Download (PDF) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Links | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| External Links: |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
Enzymes
- General function:
- Involved in carboxyl- or carbamoyltransferase activity
- Specific function:
- Carbamoyl phosphate + L-ornithine = phosphate + L-citrulline
- Gene Name:
- argF
- Locus Tag:
- PA3537
- Molecular weight:
- 33.9 kDa
Reactions
| Carbamoyl phosphate + L-ornithine = phosphate + L-citrulline. |
- General function:
- Involved in nucleotide binding
- Specific function:
- Part of the binding-protein-dependent transport system for histidine. Probably responsible for energy coupling to the transport system
- Gene Name:
- hisP
- Locus Tag:
- PA2926
- Molecular weight:
- 28.5 kDa
- General function:
- Involved in carboxy-lyase activity
- Specific function:
- L-ornithine = putrescine + CO(2)
- Gene Name:
- speC
- Locus Tag:
- PA4519
- Molecular weight:
- 43.6 kDa
Reactions
| L-ornithine = putrescine + CO(2). |
- General function:
- Involved in zinc ion binding
- Specific function:
- Displays a broad specificity and can also deacylate substrates such as acetylarginine, acetylhistidine or acetylglutamate semialdehyde
- Gene Name:
- argE
- Locus Tag:
- PA5206
- Molecular weight:
- 42.2 kDa
Reactions
| N(2)-acetyl-L-ornithine + H(2)O = acetate + L-ornithine. |
- General function:
- Involved in transporter activity
- Specific function:
- Part of the binding-protein-dependent transport system for histidine; probably responsible for the translocation of the substrate across the membrane
- Gene Name:
- hisM
- Locus Tag:
- PA2925
- Molecular weight:
- 26.7 kDa
- General function:
- Involved in transporter activity
- Specific function:
- Part of the binding-protein-dependent transport system for histidine; probably responsible for the translocation of the substrate across the membrane
- Gene Name:
- hisQ
- Locus Tag:
- PA2924
- Molecular weight:
- 24.5 kDa
Transporters
- General function:
- Involved in nucleotide binding
- Specific function:
- Part of the binding-protein-dependent transport system for histidine. Probably responsible for energy coupling to the transport system
- Gene Name:
- hisP
- Locus Tag:
- PA2926
- Molecular weight:
- 28.5 kDa
- General function:
- Involved in transporter activity
- Specific function:
- Part of the binding-protein-dependent transport system for histidine; probably responsible for the translocation of the substrate across the membrane
- Gene Name:
- hisM
- Locus Tag:
- PA2925
- Molecular weight:
- 26.7 kDa
- General function:
- Involved in transporter activity
- Specific function:
- Part of the binding-protein-dependent transport system for histidine; probably responsible for the translocation of the substrate across the membrane
- Gene Name:
- hisQ
- Locus Tag:
- PA2924
- Molecular weight:
- 24.5 kDa