|
Record Information |
|---|
| Version |
1.0 |
|---|
| Update Date |
1/22/2018 12:54:54 PM |
|---|
|
Metabolite ID | PAMDB120165 |
|---|
|
Identification |
|---|
| Name: |
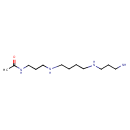
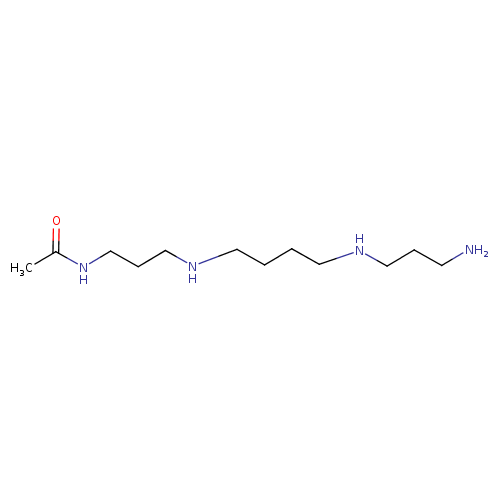
N1-acetylspermine |
|---|
| Description: | Trication of N1-acetylspermine arising from protonation of the one primary and two secondary amino groups; major species at pH 7.3. |
|---|
|
Structure |
|
|---|
| Synonyms: | - N1-acetylspermine
- N1-acetylsperminium trication
|
|---|
|
Chemical Formula: |
C12H31N4O |
|---|
| Average Molecular Weight: |
247.403 |
|---|
| Monoisotopic Molecular
Weight: |
247.24979 |
|---|
| InChI Key: |
GUNURVWAJRRUAV-UHFFFAOYSA-Q |
|---|
| InChI: | InChI=1S/C12H28N4O/c1-12(17)16-11-5-10-15-8-3-2-7-14-9-4-6-13/h14-15H,2-11,13H2,1H3,(H,16,17)/p+3 |
|---|
| CAS
number: |
25593-72-0 |
|---|
| IUPAC Name: | N-(3-acetamidopropyl)-N'-(3-azaniumylpropyl)butane-1,4-diaminium |
|---|
|
Traditional IUPAC Name: |
N(1)-acetylspermine |
|---|
| SMILES: | CC(=O)NCCC[N+]CCCC[N+]CCC[N+] |
|---|
|
Chemical Taxonomy |
|---|
|
Taxonomy Description | This compound belongs to the class of chemical entities known as acetamides. These are organic compounds with the general formula RNHC(=O)CH3, where R= organyl group. |
|---|
|
Kingdom |
Chemical entities |
|---|
| Super Class | Organic compounds |
|---|
|
Class |
Organic acids and derivatives |
|---|
| Sub Class | Carboxylic acids and derivatives |
|---|
|
Direct Parent |
Acetamides |
|---|
| Alternative Parents |
|
|---|
| Substituents |
- Acetamide
- Amino acid or derivatives
- Secondary carboxylic acid amide
- Secondary aliphatic amine
- Secondary amine
- Amine
- Hydrocarbon derivative
- Organic oxide
- Organopnictogen compound
- Primary amine
- Organooxygen compound
- Organonitrogen compound
- Primary aliphatic amine
- Organic oxygen compound
- Carbonyl group
- Organic nitrogen compound
- Aliphatic acyclic compound
|
|---|
| Molecular Framework |
Aliphatic acyclic compounds |
|---|
| External Descriptors |
|
|---|
|
Physical Properties |
|---|
| State: |
Solid |
|---|
| Charge: | 3 |
|---|
|
Melting point: |
Not Available |
|---|
| Experimental Properties: |
| Property | Value | Reference |
|---|
| Melting Point | Not Available | Not Available | | Boiling Point | Not Available | Not Available | | Water Solubility | Not Available | Not Available | | LogP | Not Available | Not Available |
|
|---|
| Predicted Properties |
|
|---|
|
Biological Properties |
|---|
| Cellular Locations: |
Not Available |
|---|
| Reactions: | |
|---|
|
Pathways: |
Not Available |
|---|
|
Spectra |
|---|
| Spectra: |
| Spectrum Type | Description | Splash Key | |
|---|
| Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS | Not Available |
|---|
| LC-MS/MS | LC-MS/MS Spectrum - Quattro_QQQ 10V, Positive (Annotated) | splash10-0097-0960000000-75020060f674e61ca7f5 | View in MoNA |
|---|
| LC-MS/MS | LC-MS/MS Spectrum - Quattro_QQQ 25V, Positive (Annotated) | splash10-01ot-9500000000-f7fc690c89dd182bd5ee | View in MoNA |
|---|
| LC-MS/MS | LC-MS/MS Spectrum - Quattro_QQQ 40V, Positive (Annotated) | splash10-001i-9100000000-1d31e6d5c8f815f68e8c | View in MoNA |
|---|
| LC-MS/MS | LC-MS/MS Spectrum - LC-ESI-QQ (API3000, Applied Biosystems) 10V, Positive | splash10-0002-0090000000-09b81054799f3b804bf7 | View in MoNA |
|---|
| LC-MS/MS | LC-MS/MS Spectrum - LC-ESI-QQ (API3000, Applied Biosystems) 20V, Positive | splash10-00ba-0920000000-661cc70e5955c3a4423e | View in MoNA |
|---|
| LC-MS/MS | LC-MS/MS Spectrum - LC-ESI-QQ (API3000, Applied Biosystems) 30V, Positive | splash10-0ik9-1900000000-99bd1a086e687629194d | View in MoNA |
|---|
| LC-MS/MS | LC-MS/MS Spectrum - LC-ESI-QQ (API3000, Applied Biosystems) 40V, Positive | splash10-0il0-5900000000-181066e0b9be9cd09644 | View in MoNA |
|---|
| LC-MS/MS | LC-MS/MS Spectrum - LC-ESI-QQ (API3000, Applied Biosystems) 50V, Positive | splash10-001i-9300000000-0ab0b3f7fb3eb674343e | View in MoNA |
|---|
| LC-MS/MS | LC-MS/MS Spectrum - LC-ESI-IT (LC/MSD Trap XCT, Agilent Technologies) , Positive | splash10-00b9-0910000000-0ded9399ebe4b3d25073 | View in MoNA |
|---|
| LC-MS/MS | LC-MS/MS Spectrum - LC-ESI-IT (LC/MSD Trap XCT, Agilent Technologies) , Positive | splash10-0udi-0900000000-54a2cb1d413b63cacb96 | View in MoNA |
|---|
| LC-MS/MS | LC-MS/MS Spectrum - LC-ESI-IT (LC/MSD Trap XCT, Agilent Technologies) , Positive | splash10-03di-0900000000-2b9ff6380c30a974293f | View in MoNA |
|---|
| LC-MS/MS | LC-MS/MS Spectrum - LC-ESI-IT (LC/MSD Trap XCT, Agilent Technologies) , Positive | splash10-0udi-1900000000-92a251b4b894d28948c1 | View in MoNA |
|---|
| LC-MS/MS | LC-MS/MS Spectrum - LC-ESI-IT (LC/MSD Trap XCT, Agilent Technologies) , Positive | splash10-03di-0900000000-4a8af7c0a23d4640e59a | View in MoNA |
|---|
| LC-MS/MS | LC-MS/MS Spectrum - LC-ESI-QTOF (UPLC Q-Tof Premier, Waters) , Positive | splash10-0wc1-1910000000-df55bc991e81a351aaaa | View in MoNA |
|---|
| Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Positive | Not Available |
|---|
| Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Positive | Not Available |
|---|
| Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Positive | Not Available |
|---|
| Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Negative | Not Available |
|---|
| Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Negative | Not Available |
|---|
| Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Negative | Not Available |
|---|
| 1D NMR | 1H NMR Spectrum | Not Available |
|---|
| 2D NMR | [1H,13C] 2D NMR Spectrum | Not Available |
|---|
|
|---|
|
References |
|---|
| References: |
- Fogel WA, Maslinski C: The FAD dependent amine oxidases in relation to developmental state of enterocyte. J Neural Transm Suppl. 1994;41:95-9. [7931271 ]
|
|---|
| Synthesis Reference: |
Not Available |
|---|
| Material Safety Data Sheet (MSDS) |
Download (PDF) |
|---|
|
Links |
|---|
| External Links: |
|
|---|