|
Record Information |
|---|
| Version |
1.0 |
|---|
| Update Date |
1/22/2018 12:54:54 PM |
|---|
|
Metabolite ID | PAMDB001449 |
|---|
|
Identification |
|---|
| Name: |
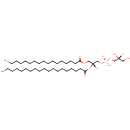
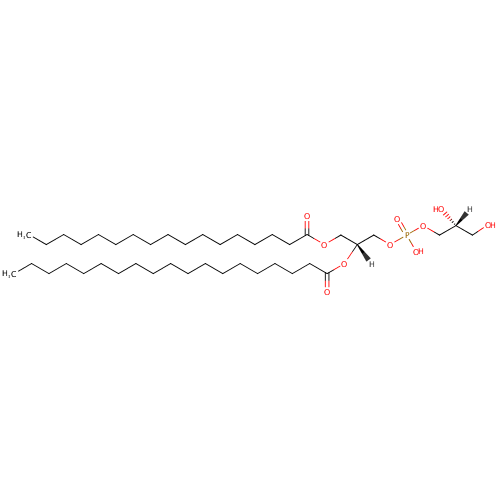
PG(19:0/17:0) |
|---|
| Description: | PG(19:0/17:0) is a phosphatidylglycerol. Phosphatidylglycerols consist of a glycerol 3-phosphate backbone esterified to either saturated or unsaturated fatty acids on carbons 1 and 2. As is the case with diacylglycerols, phosphatidylglycerols can have many different combinations of fatty acids of varying lengths and saturation attached to the C-1 and C-2 positions. PG(19:0/17:0), in particular, consists of one nonadecanoyl chain to the C-1 atom, and one heptadecanoyl to the C-2 atom. In Pseudomonas aeruginosa glycerophospholipid metabolism, phosphatidylglycerol is formed from phosphatidic acid (1,2-diacyl-sn-glycerol 3-phosphate) by a sequence of enzymatic reactions that proceeds via two intermediates, cytidine diphosphate diacylglycerol (CDP-diacylglycerol) and phosphatidylglycerophosphate (PGP, a phosphorylated phosphatidylglycerol). Phosphatidylglycerols, along with CDP-diacylglycerol, also serve as precursor molecules for the synthesis of cardiolipin, a phospholipid found in membranes. |
|---|
|
Structure |
|
|---|
| Synonyms: | - 1--2-heptadecanoyl-sn-glycero-3-phosphoglycerol
- 1-nonadecanoyl-2-heptadecanoyl-glycero-3-phospho-(1'-sn-glycerol)
- 1-nonadecanoyl-2-heptadecanoyl-sn-glycero-3-phosphoglycerol
- 1-nonadecanoyl-2-margaroyl-sn-glycero-3-phospho-(1'-glycerol)
- 1-nonadecanoyl-2-margaroyl-sn-glycero-3-phosphoglycerol
- GPG(19:0/17:0)
- GPG(36:0)
- PG(36:0)
- Phosphatidylglycerol(19:0/17:0)
- Phosphatidylglycerol(36:0)
|
|---|
|
Chemical Formula: |
C42H83O10P |
|---|
| Average Molecular Weight: |
779.09 |
|---|
| Monoisotopic Molecular
Weight: |
778.572385871 |
|---|
| InChI Key: |
VRXBINAFDUHENA-XRSDMRJBSA-N |
|---|
| InChI: | InChI=1S/C42H83O10P/c1-3-5-7-9-11-13-15-17-19-20-22-24-26-28-30-32-34-42(46)52-40(38-51-53(47,48)50-36-39(44)35-43)37-49-41(45)33-31-29-27-25-23-21-18-16-14-12-10-8-6-4-2/h39-40,43-44H,3-38H2,1-2H3,(H,47,48)/t39-,40-/m1/s1 |
|---|
| CAS
number: |
Not Available |
|---|
| IUPAC Name: | [(2R)-2,3-dihydroxypropoxy][(2R)-3-(heptadecanoyloxy)-2-(nonadecanoyloxy)propoxy]phosphinic acid |
|---|
|
Traditional IUPAC Name: |
(2R)-2,3-dihydroxypropoxy((2R)-3-(heptadecanoyloxy)-2-(nonadecanoyloxy)propoxy)phosphinic acid |
|---|
| SMILES: | [H][C@@](O)(CO)COP(O)(=O)OC[C@@]([H])(COC(=O)CCCCCCCCCCCCCCCC)OC(=O)CCCCCCCCCCCCCCCCCC |
|---|
|
Chemical Taxonomy |
|---|
|
Taxonomy Description | Not Available |
|---|
|
Kingdom |
Not Available |
|---|
| Super Class | Not Available |
|---|
|
Class |
Not Available |
|---|
| Sub Class | Not Available |
|---|
|
Direct Parent |
Not Available |
|---|
| Alternative Parents |
Not Available |
|---|
| Substituents |
Not Available |
|---|
| Molecular Framework |
Not Available |
|---|
| External Descriptors |
Not Available |
|---|
|
Physical Properties |
|---|
| State: |
Not Available |
|---|
| Charge: | -1 |
|---|
|
Melting point: |
Not Available |
|---|
| Experimental Properties: |
|
|---|
| Predicted Properties |
|
|---|
|
Biological Properties |
|---|
| Cellular Locations: |
Membrane |
|---|
| Reactions: | |
|---|
|
Pathways: |
|
|---|
|
Spectra |
|---|
| Spectra: |
|
|---|
|
References |
|---|
| References: |
- Kanehisa, M., Goto, S., Sato, Y., Furumichi, M., Tanabe, M. (2012). "KEGG for integration and interpretation of large-scale molecular data sets." Nucleic Acids Res 40:D109-D114. Pubmed: 22080510
- Keseler, I. M., Collado-Vides, J., Santos-Zavaleta, A., Peralta-Gil, M., Gama-Castro, S., Muniz-Rascado, L., Bonavides-Martinez, C., Paley, S., Krummenacker, M., Altman, T., Kaipa, P., Spaulding, A., Pacheco, J., Latendresse, M., Fulcher, C., Sarker, M., Shearer, A. G., Mackie, A., Paulsen, I., Gunsalus, R. P., Karp, P. D. (2011). "EcoCyc: a comprehensive database of Escherichia coli biology." Nucleic Acids Res 39:D583-D590. Pubmed: 21097882
- Uniprot Consortium (2012). "Reorganizing the protein space at the Universal Protein Resource (UniProt)." Nucleic Acids Res 40:D71-D75. Pubmed: 22102590
- Yurtsever D. (2007). Fatty acid methyl ester profiling of Enterococcus and Esherichia coli for microbial source tracking. M.sc. Thesis. Villanova University: U.S.A
|
|---|
| Synthesis Reference: |
Not Available |
|---|
| Material Safety Data Sheet (MSDS) |
Not Available |
|---|
|
Links |
|---|
| External Links: |
| Resource | Link |
|---|
| CHEBI ID | Not Available | | HMDB ID | Not Available | | Pubchem Compound ID | Not Available | | Kegg ID | Not Available | | ChemSpider ID | Not Available | | Wikipedia ID | Not Available | | BioCyc ID | Not Available |
|
|---|