D-Erythrose 4-phosphate (PAMDB000460)
| Record Information | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Version | 1.0 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Update Date | 1/22/2018 12:54:54 PM | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Metabolite ID | PAMDB000460 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Identification | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
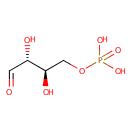
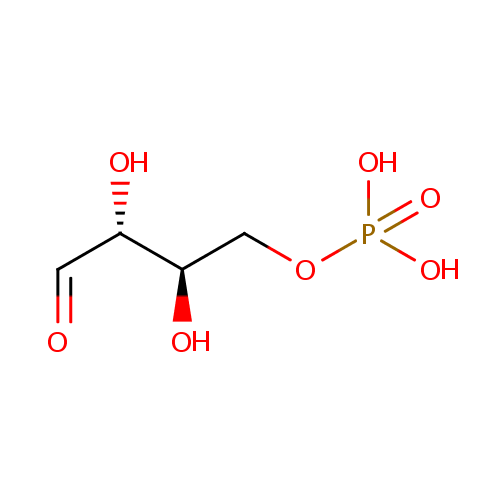
| Name: | D-Erythrose 4-phosphate | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Description: | D-Erythrose 4-phosphate is a phosphorylated derivative of erythrose that serves as an important intermediate in the pentose phosphate pathway. It is also used in phenylalanine, tyrosine and tryptophan biosynthesis, and it plays a role in vitamin B6 metabolism (KEGG) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Structure | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Synonyms: |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Chemical Formula: | C4H9O7P | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Average Molecular Weight: | 200.0838 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Monoisotopic Molecular Weight: | 200.008589154 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| InChI Key: | NGHMDNPXVRFFGS-IUYQGCFVSA-N | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| InChI: | InChI=1S/C4H9O7P/c5-1-3(6)4(7)2-11-12(8,9)10/h1,3-4,6-7H,2H2,(H2,8,9,10)/t3-,4+/m0/s1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| CAS number: | 585-18-2 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| IUPAC Name: | [(2R,3R)-2,3-dihydroxy-4-oxobutoxy]phosphonic acid | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Traditional IUPAC Name: | 4-O-phosphono-D-erythrose | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| SMILES: | O[C@H](COP(O)(O)=O)[C@@H](O)C=O | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Chemical Taxonomy | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Taxonomy Description | This compound belongs to the class of organic compounds known as monosaccharide phosphates. These are monosaccharides comprising a phosphated group linked to the carbohydrate unit. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Kingdom | Organic compounds | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Super Class | Organooxygen compounds | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Class | Carbohydrates and carbohydrate conjugates | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Sub Class | Monosaccharides | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Direct Parent | Monosaccharide phosphates | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Alternative Parents | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Substituents |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Molecular Framework | Aliphatic acyclic compounds | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| External Descriptors |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Physical Properties | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| State: | Solid | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Charge: | -2 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Melting point: | Not Available | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Experimental Properties: |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Predicted Properties |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Biological Properties | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Cellular Locations: | Cytoplasm | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Reactions: | D-Glyceraldehyde 3-phosphate + D-Sedoheptulose 7-phosphate <> D-Erythrose 4-phosphate + Fructose 6-phosphate D-Erythrose 4-phosphate + Water + Phosphoenolpyruvic acid <> 2-Dehydro-3-deoxy-D-arabino-heptonate 7-phosphate + Phosphate D-Erythrose 4-phosphate + Water + NAD <> 4-Phospho-D-erythronate +2 Hydrogen ion + NADH Sedoheptulose 1,7-bisphosphate <> Dihydroxyacetone phosphate + D-Erythrose 4-phosphate Sedoheptulose 7-phosphate + D-Glyceraldehyde 3-phosphate <> D-Erythrose 4-phosphate + beta-D-Fructose 6-phosphate beta-D-Fructose 6-phosphate + D-Glyceraldehyde 3-phosphate <> D-Erythrose 4-phosphate + Xylulose 5-phosphate D-Erythrose 4-phosphate + Xylulose 5-phosphate <> Fructose 6-phosphate + D-Glyceraldehyde 3-phosphate Phosphoenolpyruvic acid + D-Erythrose 4-phosphate + Water > 2-Dehydro-3-deoxy-D-arabino-heptonate 7-phosphate + Inorganic phosphate D-Erythrose 4-phosphate + NAD + Water > 4-Phospho-D-erythronate + NADH D-Sedoheptulose 7-phosphate + D-Glyceraldehyde 3-phosphate + D-Sedoheptulose 7-phosphate + D-Glyceraldehyde 3-phosphate <> beta-D-Fructose 6-phosphate + D-Erythrose 4-phosphate D-Erythrose 4-phosphate + Water + Phosphoenolpyruvic acid > Phosphate + 3-deoxy-D-arabino-heptulosonate-7-phosphate | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Pathways: | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Spectra | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Spectra: |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| References | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| References: |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Synthesis Reference: | Not Available | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Material Safety Data Sheet (MSDS) | Not Available | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Links | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| External Links: |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||