5'-Phosphoribosyl-N-formylglycineamide (PAMDB000332)
| Record Information | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Version | 1.0 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Update Date | 1/22/2018 12:54:54 PM | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Metabolite ID | PAMDB000332 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Identification | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
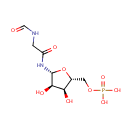
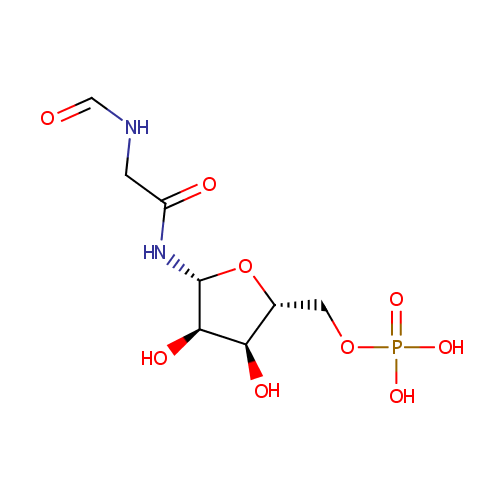
| Name: | 5'-Phosphoribosyl-N-formylglycineamide | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Description: | 5'-Phosphoribosyl-N-formylglycineamide (also known as FGAR or N-Formyl-GAR) is a substrate for Phosphoribosylformylglycinamidine synthase. It is involved in aminoimidazole ribonucleotide biosynthesis and plays a vital role in purine metabolism as well as the conversion of glutamine to glutamate. 5'-Phosphoribosyl-N-formylglycineamide is described as a glycinamide ribonucleotide having a phosphate group at the 5-position and a formyl group on the glycine nitrogen. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Structure | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Synonyms: |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Chemical Formula: | C8H15N2O9P | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Average Molecular Weight: | 314.1865 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Monoisotopic Molecular Weight: | 314.0515166 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| InChI Key: | VDXLUNDMVKSKHO-XVFCMESISA-N | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| InChI: | InChI=1S/C8H15N2O9P/c11-3-9-1-5(12)10-8-7(14)6(13)4(19-8)2-18-20(15,16)17/h3-4,6-8,13-14H,1-2H2,(H,9,11)(H,10,12)(H2,15,16,17)/t4-,6-,7-,8-/m1/s1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| CAS number: | Not Available | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| IUPAC Name: | {[(2R,3S,4R,5R)-3,4-dihydroxy-5-(2-formamidoacetamido)oxolan-2-yl]methoxy}phosphonic acid | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Traditional IUPAC Name: | [(2R,3S,4R,5R)-3,4-dihydroxy-5-(2-formamidoacetamido)oxolan-2-yl]methoxyphosphonic acid | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| SMILES: | O[C@H]1[C@@H](O)[C@H](NC(=O)CNC=O)O[C@@H]1COP(O)(O)=O | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Chemical Taxonomy | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Taxonomy Description | This compound belongs to the class of organic compounds known as glycinamide ribonucleotides. These are compounds in which the?amide?N atom of glycineamide is?linked?to the C-1 of a?ribosyl (or deoxyribosyl) moiety. Nucleotides have a phosphate group linked to the C5 carbon of the ribose (or deoxyribose) moiety. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Kingdom | Organic compounds | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Super Class | Nucleosides, nucleotides, and analogues | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Class | Glycinamide ribonucleotides | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Sub Class | Not Available | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Direct Parent | Glycinamide ribonucleotides | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Alternative Parents | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Substituents |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Molecular Framework | Aliphatic heteromonocyclic compounds | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| External Descriptors | Not Available | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Physical Properties | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| State: | Not Available | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Charge: | -2 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Melting point: | Not Available | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Experimental Properties: |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Predicted Properties |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Biological Properties | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Cellular Locations: | Cytoplasm | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Reactions: | Adenosine triphosphate + Formic acid + Glycineamideribotide > ADP + 5'-Phosphoribosyl-N-formylglycineamide + Hydrogen ion + Phosphate N10-Formyl-THF + Glycineamideribotide <> 5'-Phosphoribosyl-N-formylglycineamide + Hydrogen ion + Tetrahydrofolic acid Adenosine triphosphate + 5'-Phosphoribosyl-N-formylglycineamide + L-Glutamine + Water <> ADP + Phosphoribosylformylglycineamidine + L-Glutamate + Hydrogen ion + Phosphate N10-Formyl-THF + Glycineamideribotide <> Tetrahydrofolic acid + 5'-Phosphoribosyl-N-formylglycineamide Glycineamideribotide + 5,10-Methenyltetrahydrofolate + Water <> 5'-Phosphoribosyl-N-formylglycineamide + Tetrahydrofolic acid Adenosine triphosphate + 5'-Phosphoribosyl-N-formylglycineamide + L-Glutamine + Water <> ADP + Phosphate + Phosphoribosylformylglycineamidine + L-Glutamate Formic acid + Adenosine triphosphate + 5'-Phospho-ribosylglycinamide > 5'-Phosphoribosyl-N-formylglycineamide + ADP + Pyrophosphate Tetrahydrofolic acid + 5'-Phosphoribosyl-N-formylglycinamide + Tetrahydrofolic acid + 5'-Phosphoribosyl-N-formylglycineamide > Water + 5,10-Methenyltetrahydrofolic acid + Glycineamideribotide + Glycineamideribotide Tetrahydrofolic acid + 5'-Phosphoribosyl-N-formylglycinamide + Tetrahydrofolic acid + 5'-Phosphoribosyl-N-formylglycineamide > 10-Formyltetrahydrofolate + Glycineamideribotide + N10-Formyl-THF + Glycineamideribotide N1-(5-phospho-β-D-ribosyl)glycinamide + Adenosine triphosphate + Formic acid > 5'-Phosphoribosyl-N-formylglycinamide + Adenosine diphosphate + Phosphate + Hydrogen ion + 5'-Phosphoribosyl-N-formylglycineamide + ADP 5'-Phosphoribosyl-N-formylglycinamide + Water + L-Glutamine + Adenosine triphosphate + 5'-Phosphoribosyl-N-formylglycineamide > 2-(Formamido)-N1-(5-phospho-D-ribosyl)acetamidine + L-Glutamic acid + Phosphate + Adenosine diphosphate + Hydrogen ion + L-Glutamate + ADP | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Pathways: | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Spectra | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Spectra: |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| References | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| References: |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Synthesis Reference: | Not Available | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Material Safety Data Sheet (MSDS) | Not Available | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Links | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| External Links: |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||