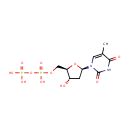
dTDP (PAMDB000320)
| Record Information | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Version | 1.0 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Update Date | 1/22/2018 12:54:54 PM | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Metabolite ID | PAMDB000320 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Identification | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Name: | dTDP | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Description: | dTDP is an intermediate in the synthesis and breakdown of DNA. Thymidylate kinase (EC 2.7.4.9; ATP:dTMP phosphotransferase) catalyzes the phosphorylation of dTMP (to form dTDP) in the dTTP synthesis pathway for DNA synthesis. (OMIM 188345 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Structure | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Synonyms: |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Chemical Formula: | C10H16N2O11P2 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Average Molecular Weight: | 402.1884 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Monoisotopic Molecular Weight: | 402.022932388 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| InChI Key: | UJLXYODCHAELLY-XLPZGREQSA-N | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| InChI: | InChI=1S/C10H16N2O11P2/c1-5-3-12(10(15)11-9(5)14)8-2-6(13)7(22-8)4-21-25(19,20)23-24(16,17)18/h3,6-8,13H,2,4H2,1H3,(H,19,20)(H,11,14,15)(H2,16,17,18)/t6-,7+,8+/m0/s1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| CAS number: | 491-97-4 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| IUPAC Name: | {[hydroxy({[(2R,3S,5R)-3-hydroxy-5-(5-methyl-2,4-dioxo-1,2,3,4-tetrahydropyrimidin-1-yl)oxolan-2-yl]methoxy})phosphoryl]oxy}phosphonic acid | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Traditional IUPAC Name: | dTDP | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| SMILES: | CC1=CN([C@H]2C[C@H](O)[C@@H](COP(O)(=O)OP(O)(O)=O)O2)C(=O)NC1=O | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Chemical Taxonomy | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Taxonomy Description | This compound belongs to the class of organic compounds known as pyrimidine 2'-deoxyribonucleoside diphosphates. These are pyrimidine nucleotides with a diphosphate group linked to the ribose moiety lacking a hydroxyl group at position 2. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Kingdom | Organic compounds | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Super Class | Nucleosides, nucleotides, and analogues | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Class | Pyrimidine nucleotides | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Sub Class | Pyrimidine deoxyribonucleotides | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Direct Parent | Pyrimidine 2'-deoxyribonucleoside diphosphates | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Alternative Parents | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Substituents |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Molecular Framework | Aromatic heteromonocyclic compounds | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| External Descriptors |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Physical Properties | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| State: | Solid | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Charge: | -2 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Melting point: | Not Available | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Experimental Properties: |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Predicted Properties |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Biological Properties | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Cellular Locations: | Cytoplasm | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Reactions: | Adenosine triphosphate + dTDP <> ADP + Thymidine 5'-triphosphate Adenosine triphosphate + 5-Thymidylic acid <> ADP + dTDP Deoxythymidine diphosphate-L-rhamnose + Kdo-phospho-heptosyl-phospho-heptosyl-heptosyl-kdo2-lipidA > dTDP + Hydrogen ion + inner core oligosaccharide lipid A (E coli) dTDP-4-Acetamido-4,6-dideoxy-D-galactose + Undecaprenyl-diphospho-N-acetylglucosamine-N-acetylmannosaminuronate > dTDP + Hydrogen ion + Undecaprenyl-diphospho N-acetylglucosamine-N-acetylmannosaminuronate-N-acetamido-4,6-dideoxy-D-galactose Thymidine 5'-triphosphate + Cytidine <> dTDP + Cytidine monophosphate Thymidine 5'-triphosphate + Uridine <> dTDP + Uridine 5'-monophosphate a lipopolysaccharide + Deoxythymidine diphosphate-L-rhamnose a rhamnosyl lipopolysaccharide + dTDP Adenosine triphosphate + dTDP > Adenosine diphosphate + Thymidine 5'-triphosphate + ADP Undecaprenyl N-acetyl-glucosaminyl-N-acetyl-mannosaminuronate pyrophosphate + TDP-Fuc4NAc + Undecaprenyl-diphospho-N-acetylglucosamine-N-acetylmannosaminuronate > Hydrogen ion + dTDP + Undecaprenyl N-acetyl-glucosaminyl-N-acetyl-mannosaminuronate-4-acetamido-4,6-dideoxy-D-galactose pyrophosphate + Undecaprenyl-diphospho N-acetylglucosamine-N-acetylmannosaminuronate-N-acetamido-4,6-dideoxy-D-galactose | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Pathways: | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Spectra | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Spectra: |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| References | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| References: |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Synthesis Reference: | Rupprath, Carsten; Kopp, Maren; Hirtz, Dennis; Mueller, Rolf; Elling, Lothar. An enzyme module system for in situ regeneration of deoxythymidine 5'-diphosphate (dTDP)-activated deoxy sugars. Advanced Synthesis & Catalysis (2007), 349(8+9), 1489-149 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Material Safety Data Sheet (MSDS) | Not Available | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Links | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| External Links: |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
Enzymes
- General function:
- Involved in thymidylate kinase activity
- Specific function:
- Catalyzes the reversible phosphorylation of deoxythymidine monophosphate (dTMP) to deoxythymidine diphosphate (dTDP), using ATP as its preferred phosphoryl donor. Situated at the junction of both de novo and salvage pathways of deoxythymidine triphosphate (dTTP) synthesis, is essential for DNA synthesis and cellular growth
- Gene Name:
- tmk
- Locus Tag:
- PA2962
- Molecular weight:
- 23.1 kDa
Reactions
| ATP + dTMP = ADP + dTDP. |
- General function:
- Involved in nucleoside diphosphate kinase activity
- Specific function:
- Major role in the synthesis of nucleoside triphosphates other than ATP. The ATP gamma phosphate is transferred to the NDP beta phosphate via a ping-pong mechanism, using a phosphorylated active-site intermediate
- Gene Name:
- ndk
- Locus Tag:
- PA3807
- Molecular weight:
- 15.6 kDa
Reactions
| ATP + nucleoside diphosphate = ADP + nucleoside triphosphate. |
- General function:
- Involved in ATP binding
- Specific function:
- Catalyzes the reversible transfer of the terminal phosphate group between ATP and AMP. This small ubiquitous enzyme involved in the energy metabolism and nucleotide synthesis, is essential for maintenance and cell growth
- Gene Name:
- adk
- Locus Tag:
- PA3686
- Molecular weight:
- 23.1 kDa
Reactions
| ATP + AMP = 2 ADP. |