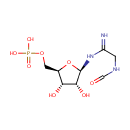
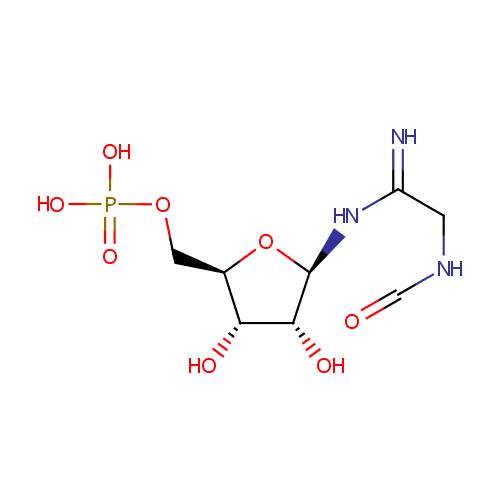
| InChI: | InChI=1S/C8H16N3O8P/c9-5(1-10-3-12)11-8-7(14)6(13)4(19-8)2-18-20(15,16)17/h3-4,6-8,13-14H,1-2H2,(H2,9,11)(H,10,12)(H2,15,16,17)/t4-,6-,7-,8-/m1/s1 |
|---|
| References: |
- Jayaram HN, Lui MS, Plowman J, Pillwein K, Reardon MA, Elliott WL, Weber G: Oncolytic activity and mechanism of action of a novel L-cysteine derivative, L-cysteine, ethyl ester, S-(N-methylcarbamate) monohydrochloride. Cancer Chemother Pharmacol. 1990;26(2):88-92. Pubmed: 2347042
- Kanehisa, M., Goto, S., Sato, Y., Furumichi, M., Tanabe, M. (2012). "KEGG for integration and interpretation of large-scale molecular data sets." Nucleic Acids Res 40:D109-D114. Pubmed: 22080510
- Keseler, I. M., Collado-Vides, J., Santos-Zavaleta, A., Peralta-Gil, M., Gama-Castro, S., Muniz-Rascado, L., Bonavides-Martinez, C., Paley, S., Krummenacker, M., Altman, T., Kaipa, P., Spaulding, A., Pacheco, J., Latendresse, M., Fulcher, C., Sarker, M., Shearer, A. G., Mackie, A., Paulsen, I., Gunsalus, R. P., Karp, P. D. (2011). "EcoCyc: a comprehensive database of Escherichia coli biology." Nucleic Acids Res 39:D583-D590. Pubmed: 21097882
- Maegawa T, Karasawa T, Ohta T, Wang X, Kato H, Hayashi H, Nakamura S: Linkage between toxin production and purine biosynthesis in Clostridium difficile. J Med Microbiol. 2002 Jan;51(1):34-41. Pubmed: 11800470
- van der Werf, M. J., Overkamp, K. M., Muilwijk, B., Coulier, L., Hankemeier, T. (2007). "Microbial metabolomics: toward a platform with full metabolome coverage." Anal Biochem 370:17-25. Pubmed: 17765195
- Winder, C. L., Dunn, W. B., Schuler, S., Broadhurst, D., Jarvis, R., Stephens, G. M., Goodacre, R. (2008). "Global metabolic profiling of Escherichia coli cultures: an evaluation of methods for quenching and extraction of intracellular metabolites." Anal Chem 80:2939-2948. Pubmed: 18331064
|
|---|