Adenine (PAMDB000010)
| Record Information | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Version | 1.0 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Update Date | 1/22/2018 12:54:54 PM | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Metabolite ID | PAMDB000010 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Identification | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
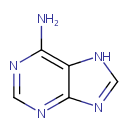
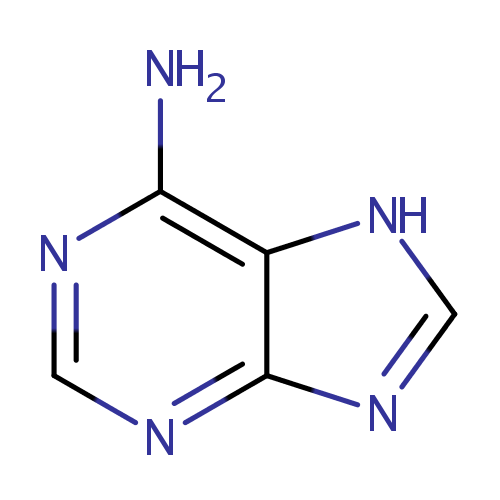
| Name: | Adenine | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Description: | Adenine is a purine base. Adenine is found in both DNA and RNA. Adenine is a fundamental component of adenine nucleotides. Adenine forms adenosine, a nucleoside, when attached to ribose, and deoxyadenosine when attached to deoxyribose; it forms adenosine triphosphate (ATP), a nucleotide, when three phosphate groups are added to adenosine. Adenosine triphosphate is used in cellular metabolism as one of the basic methods of transferring chemical energy between chemical reactions. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Structure | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Synonyms: |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Chemical Formula: | C5H5N5 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Average Molecular Weight: | 135.1267 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Monoisotopic Molecular Weight: | 135.054495185 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| InChI Key: | GFFGJBXGBJISGV-UHFFFAOYSA-N | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| InChI: | InChI=1S/C5H5N5/c6-4-3-5(9-1-7-3)10-2-8-4/h1-2H,(H3,6,7,8,9,10) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| CAS number: | 73-24-5 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| IUPAC Name: | 7H-purin-6-amine | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Traditional IUPAC Name: | vitamin B4 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| SMILES: | NC1=C2NC=NC2=NC=N1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Chemical Taxonomy | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Taxonomy Description | This compound belongs to the class of organic compounds known as 6-aminopurines. These are purines that carry an amino group at position 6. Purine is a bicyclic aromatic compound made up of a pyrimidine ring fused to an imidazole ring. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Kingdom | Organic compounds | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Super Class | Organoheterocyclic compounds | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Class | Imidazopyrimidines | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Sub Class | Purines and purine derivatives | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Direct Parent | 6-aminopurines | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Alternative Parents | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Substituents |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Molecular Framework | Aromatic heteropolycyclic compounds | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| External Descriptors |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Physical Properties | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| State: | Solid | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Charge: | 0 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Melting point: | 360 °C | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Experimental Properties: |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Predicted Properties |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Biological Properties | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Cellular Locations: | Cytoplasm | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Reactions: | Adenosine + Water > Adenine + Ribose S-Adenosylhomocysteine + Water <> Adenine + S-Ribosyl-L-homocysteine 5'-Methylthioadenosine + Water > 5-Methylthioribose + Adenine 5'-Deoxyadenosine + Water > 5'-Deoxyribose + Adenine Adenine + Phosphoribosyl pyrophosphate <> Adenosine monophosphate + Pyrophosphate Adenosine triphosphate + Dephospho-CoA > 2'-(5-Triphosphoribosyl)-3'-dephospho-CoA + Adenine Adenosine monophosphate + Water <> Adenine + D-Ribose-5-phosphate Adenine + Hydrogen ion + Water > Hypoxanthine + Ammonium Adenosine + Phosphate <> Adenine + Ribose-1-phosphate Deoxyadenosine + Phosphate <> Deoxyribose 1-phosphate + Adenine Adenine + Water <> Hypoxanthine + Ammonia Adenosine + Phosphate <> Adenine + alpha-D-Ribose 1-phosphate Methylphosphonate + Adenosine triphosphate > α-D-ribose-1-methylphosphonate-5-triphosphate + Adenine Hydrogen ion + Dephospho-CoA + Adenosine triphosphate > 2'-(5-Triphosphoribosyl)-3'-dephospho-CoA + Adenine Adenosine + Water > D-ribose + Adenine Deoxyadenosine + Phosphate <> Adenine + deoxyribose-1-phosphate More...N1-Methyladenine + Oxygen + Oxoglutaric acid > Hydrogen ion + Adenine + Carbon dioxide + Formaldehyde + Succinic acid 1-Ethyladenine + Oxygen + Oxoglutaric acid > Adenine + Carbon dioxide + Acetaldehyde + Succinic acid Adenosine triphosphate + Methylphosphonate <> alpha-D-Ribose 1-methylphosphonate 5-triphosphate + Adenine S-Adenosylhomocysteine + Water > Adenine + S-ribosyl-L-homocysteine + S-ribosyl-L-homocysteine 5'-S-methyl-5'-thioadenosine + Water > 5-Methylthioribose + Adenine 7-aminomethyl-7-deazaguanosine34 in tRNA + S-adenosyl-L-methionine > Hydrogen ion + L-Methionine + Adenine + epoxyqueuosine | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Pathways: | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Spectra | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Spectra: | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| References | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| References: |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Synthesis Reference: | Baddiley, J.; Lythgoe, B.; Todd, A. R. Synthesis of purine nucleosides. II. A new and convenient synthesis of adenine. Journal of the Chemical Society (1943), 386-7. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Material Safety Data Sheet (MSDS) | Download (PDF) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Links | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| External Links: |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||